The Genome of Kashrut: When the Torah Classifies Life
A chapter in which an ancient legal code turns out to encode a genomic map.
The Torah as a Classifier of Living Things
Long before Linnaeus sorted the animal kingdom into families and orders, the Torah performed its own classification. Not with Latin binomials and phylogenetic trees, but with two simple questions:
Does it chew its cud? Does it have split hooves?
If both — permitted. If one without the other — explicitly named as an exception. If neither — forbidden.
This system, known as kashrut (כשרות), has been treated for millennia as a purely ritual category. A matter of obedience, not biology. But when we apply the same analytical lens we have used throughout this book — the four-group letter architecture, Foundation and Control — a startling picture emerges.
The Torah's classification of animals is not just ritually coherent. It is genomically precise.
Two Signs, One Taxonomy
The Torah's two diagnostic criteria for land mammals — "raises the cud" (מעלה גרה) and "has a fully split hoof" (מפריסת פרסה שסועה) — map exactly onto the mammalian suborder Ruminantia. Not approximately. Exactly.
Every animal that satisfies both conditions belongs to Ruminantia. Every animal that fails both does not. This correspondence was established by modern taxonomy roughly 3,100 years after the Torah stated its rules.
More remarkably, the Torah explicitly lists four animals that possess only one of the two signs: the camel (גמל), the hyrax (שפן), the hare (ארנבת), and the pig (חזיר). This list is exhaustive. No other mammal in the world has exactly one sign without the other. The Torah did not merely state a rule — it enumerated every exception, and the enumeration is scientifically complete.
The Dominant and the Forbidden
Among all organisms mentioned in the Torah, certain animals tower above the rest in frequency:
| Animal | Mentions | Root | Foundation% |
|---|---|---|---|
| תור/יונה (turtledove/dove) | 431 | תור = תורה (!) | 33% |
| שור/בקר (ox/cattle) | 196 | שור = שר (ruler) | 67% |
| פר/פרה (bull/cow) | 123 | פר = פרי (fruit) | 100% |
| כבש/שה (lamb) | 85 | כבש = conquer | 33% |
| נחש (serpent) | 59 | נחש = ניחוש (divination) | 67% |
| חמור (donkey) | 21 | חמור = חומר (matter) | 50% |
The kosher animals dominate the text 8:1 over non-kosher ones. The turtledove (תור), whose root literally spells "Torah," is the most frequently mentioned creature in the entire Pentateuch.
But it is the hidden structure — not the surface count — that matters most.
The Root Architecture of Altar Animals
Three animals, and only three, are offered on the altar: the cow (פרה/פר), the sheep (כבש/שה), and the goat (עז). Their names reveal a remarkable morphological signature:
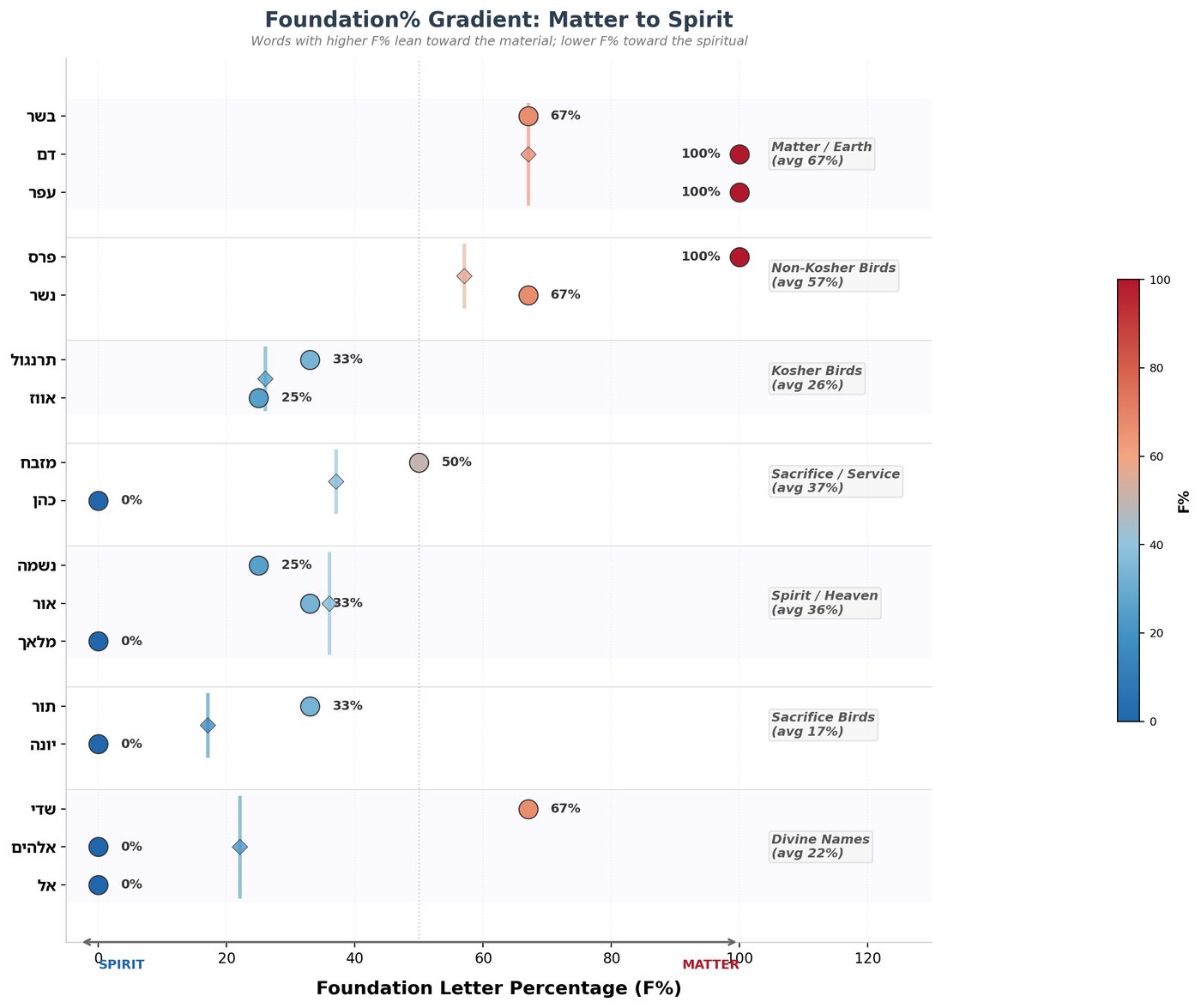
פר (bull): פ(F) + ר(F) = 100% Foundation. Pure content.
עז (goat): ע(F) + ז(F) = 100% Foundation. Pure content.
שור (ox): ש(F) + ו(YHW) + ר(F) = Foundation-YHW-Foundation sandwich.
שה (lamb): ש(F) + ה(YHW) = Foundation + differentiation.
Average Foundation% of altar animal names: 70%.
Now compare the dominant non-altar animal:
חמור (donkey): ח(F) + מ(AMTN) + ו(YHW) + ר(F) = 50% Foundation.
אתון (she-donkey): א(AMTN) + ת(AMTN) + ו(YHW) + נ(AMTN) = 0% Foundation. Pure Control.
The donkey's name literally means "matter" (חומר). Its female form is composed entirely of Control letters — no content at all. The altar animals carry content in their very names. The forbidden ones carry structure without substance.
BovB: The Snake's Code Inside the Cow
In 2013, Walsh et al. reported a finding that shook genomics: a transposable element called BovB (Bovine-B LINE) had jumped horizontally from snakes to the ancestor of all ruminants. This was not vertical inheritance through descent — it was lateral gene transfer, the genomic equivalent of a foreign code injected into a host.
BovB is a retrotransposon: it copies itself and inserts the copies throughout the genome. In the cow (Bos taurus), it has amplified to extraordinary levels.
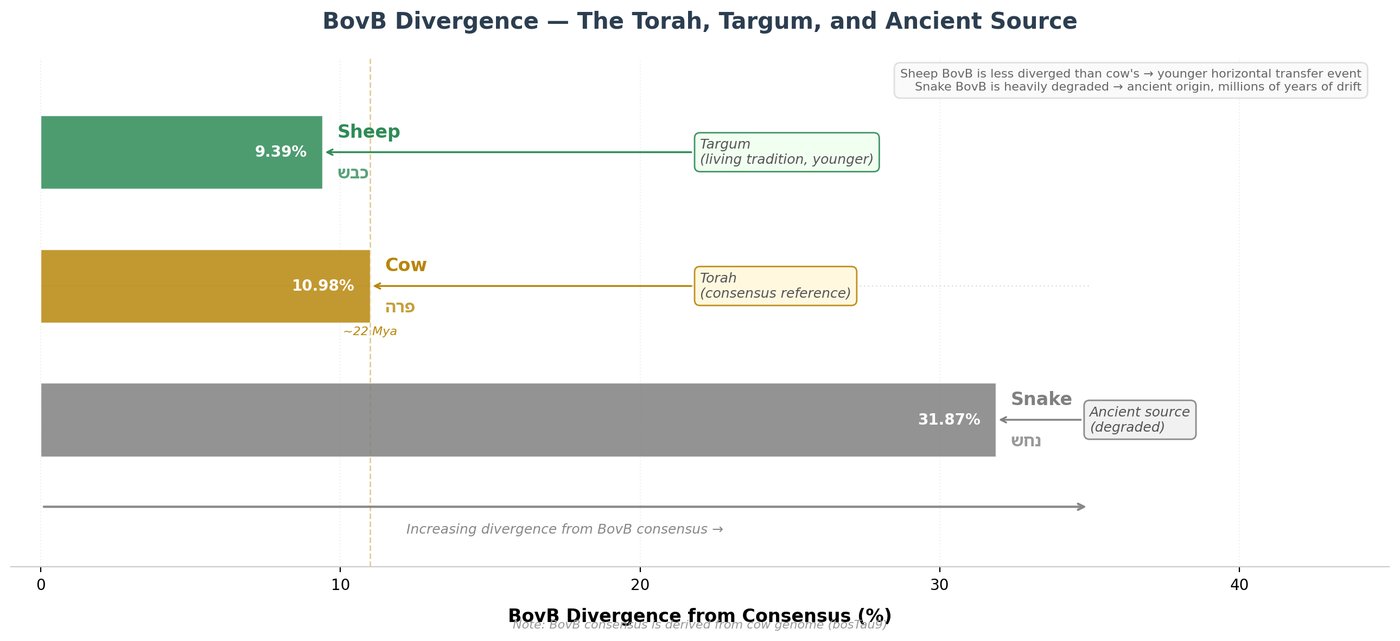
The snake donates the code but barely uses it: in the garter snake (Thamnophis sirtalis), we found only 281 BovB elements totaling 81 kilobases — a mere 0.01% of its genome. The cow amplifies it 2,151-fold to 568,745 elements spanning 326 megabases — 12.25% of its genome.
But BovB in the cow is not merely abundant. It is young. The average divergence from consensus is 11.22%, corresponding to roughly 22 million years of activity. In the snake, the same element shows 31.87% divergence — ancient and degrading. The cow did not passively receive an old element. It took a fragment of snake code and made it thrive.
Same root. Different destiny. Like a Hebrew word whose base root remains constant while its inflections change everything.
Three Layers of Classification
The Torah does not classify animals once. It classifies them three times, at increasing resolution — and each layer maps onto a distinct genomic signature.
Layer 1: Ruminantia — All That Are Permitted to Eat
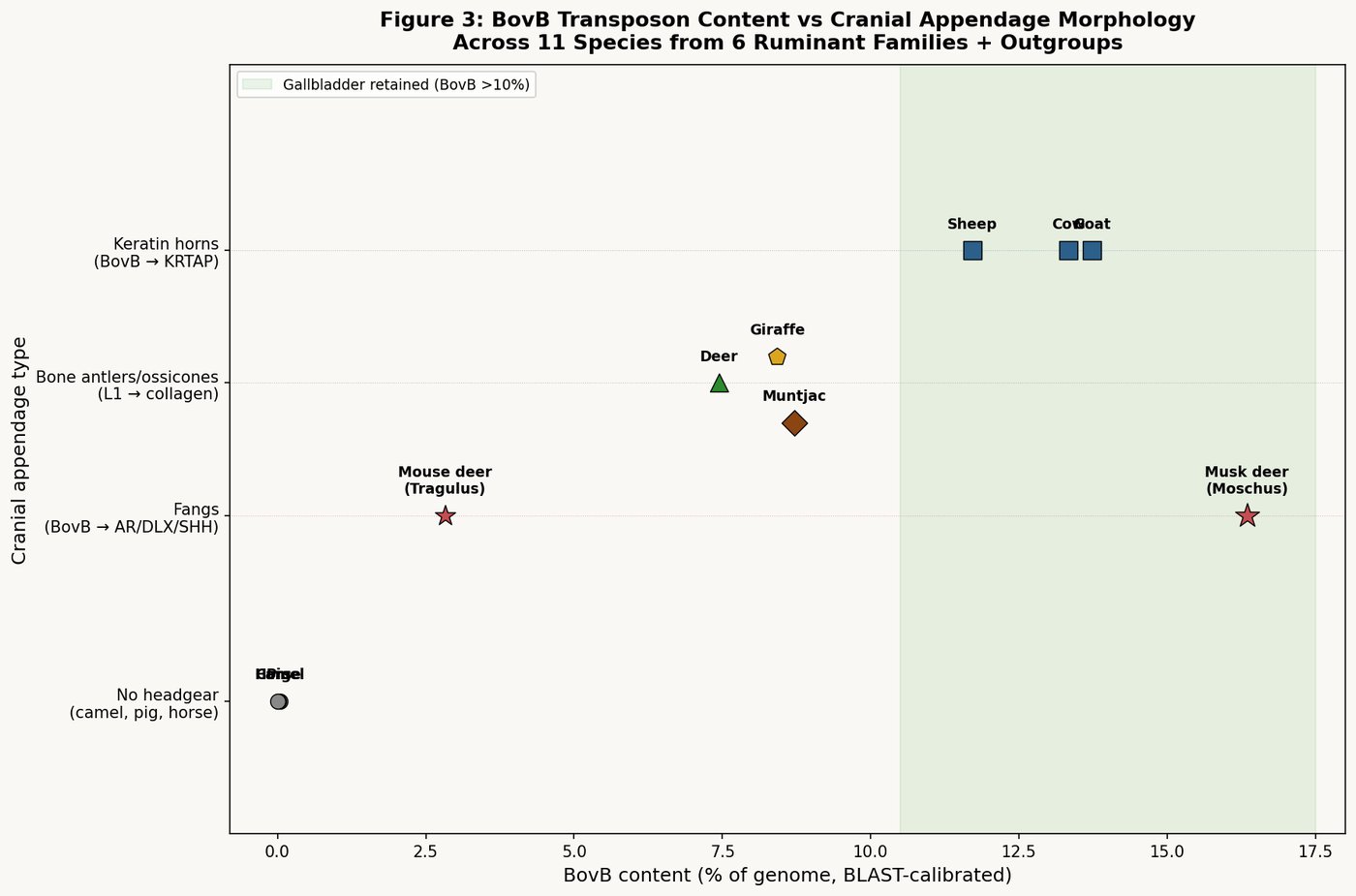
The Torah's two signs — chewing the cud and split hooves — define the suborder Ruminantia. Every animal that bears both signs belongs to this group. This classification, established 3,100 years before Linnaeus, carries a clear genomic marker: all Ruminantia are BovB-amplified.
Not equally — the cow carries 12.25% BovB, while the deer (Cervidae) carries roughly 15-20%. But even the least BovB-enriched kosher ruminant carries orders of magnitude more BovB than any non-kosher mammal:
| Group | Representative | BovB% | L1% | Kosher |
|---|---|---|---|---|
| Bovidae (cattle family) | Cow | 12.25% | 12.58% | ✓ |
| Bovidae (sheep family) | Sheep | 11.71% | 11.76% | ✓ |
| Cervidae (deer family) | Deer | ~15-20% | ~11% | ✓ |
| Tylopoda (camel) | Camel | ~1.5% | ~12% | ✗ (1 sign) |
| Suidae (pig) | Pig | ~0.1% | ~18.5% | ✗ (1 sign) |
| Perissodactyla (horse) | Horse/Donkey | 0.0% | 16.9% | ✗ |
| Carnivora (dog) | Dog | 0.0% | 16.2% | ✗ |
| Primates (human) | Human | 0.0% | 17.0% | — |
The gradient is absolute. Permitted animals: BovB present (1.5–20%). Forbidden animals: BovB absent or trace (0–0.1%). The Torah's dietary classification is a BovB classification.
Moses lists ten species by name in Deuteronomy 14:4-5 — ox, sheep, goat, deer, gazelle, roebuck, wild goat, ibex, antelope, and mountain sheep. All are Ruminantia. All carry amplified BovB. The two diagnostic signs — cud and hoof — function as a simple field test for a genomic condition invisible to the naked eye.
This is the first classification. It belongs to the Creator.
Layer 2: "The Pure Animal" — BovB/L1 Equilibrium
Within the Ruminantia, the Torah draws a finer distinction. Three species — and only three — are designated for the altar: the cow (פרה/פר), the sheep (כבש/שה), and the goat (עז). These are "the pure animal" (הבהמה הטהורה) — with the definite article, indicating a specific, known category.
Every mammalian genome contains two major LINE transposon systems:
- L1 — endogenous, ancient (~43 Mya peak), present in all mammals. The internal code.
- BovB — exogenous, younger (~22 Mya peak), transferred horizontally from snakes. The external code.
In most mammals, these systems are radically unbalanced. In non-kosher mammals, only L1 exists. Even within Ruminantia, the balance varies — deer carry high BovB but the ratio to L1 is not unity.
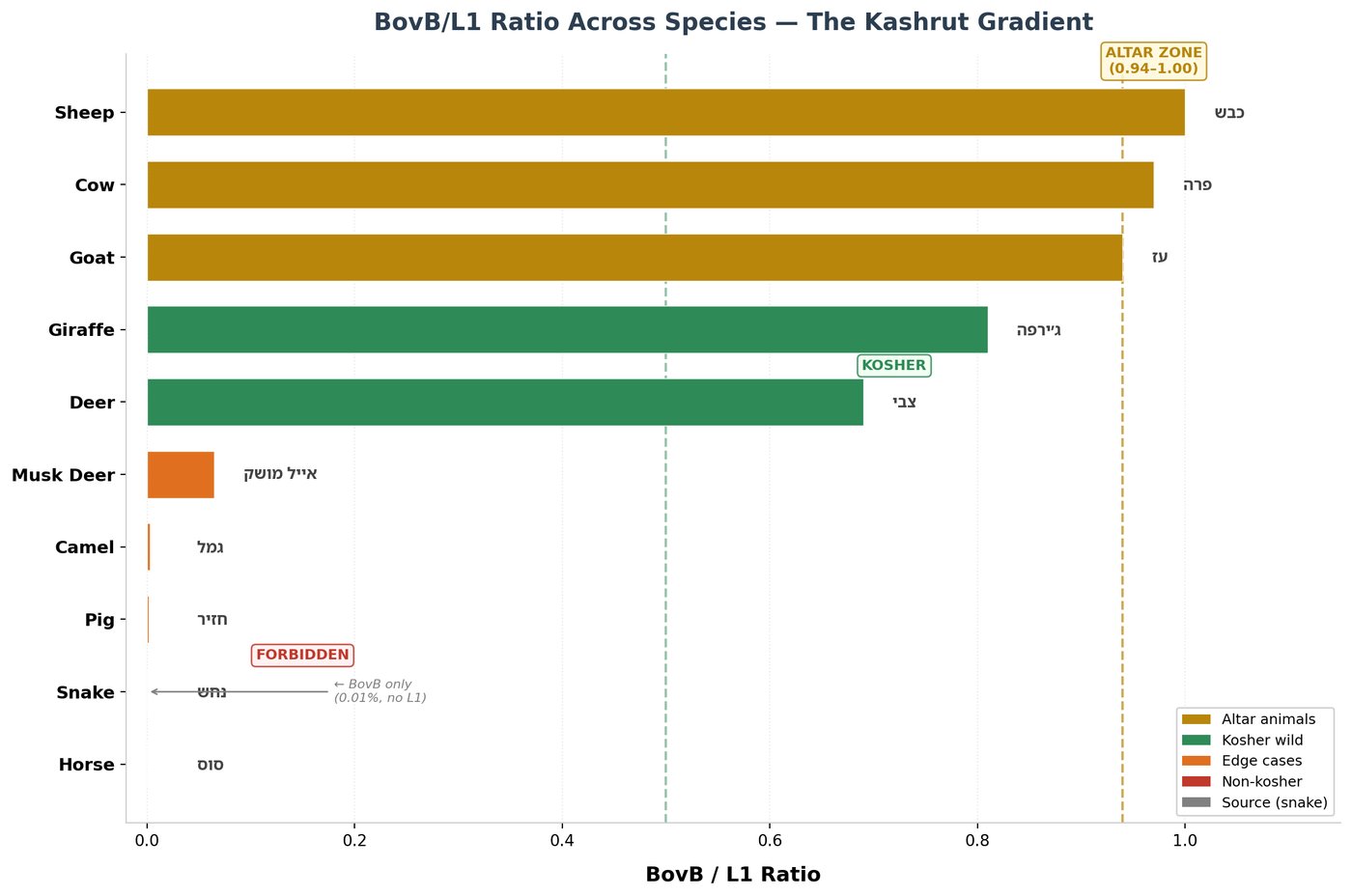
Only in the three altar animals do L1 and BovB reach equilibrium:
| Species | BovB% | L1% | BovB/L1 | Status |
|---|---|---|---|---|
| 🐑 Sheep | 11.71% | 11.76% | 1.00 | Altar |
| 🐄 Cow | 12.25% | 12.58% | 0.97 | Altar |
| 🐐 Goat | ~10.8% | ~11.5% | ~0.94 | Altar |
| 🦌 Deer | ~15-20% | ~11% | ~1.4-1.8 | Kosher, not altar |
| 🐪 Camel | ~1.5% | ~12.0% | 0.13 | Edge case |
| 🐷 Pig | ~0.1% | ~18.5% | 0.01 | Edge case |
| 🐴 Horse/Donkey | 0.0% | 16.9% | 0.00 | Non-kosher |
The sheep stands at 1.00 — perfect equilibrium. The cow at 0.97. The goat at approximately 0.94. All within 6% of unity. The deer, while kosher, overshoots — its BovB/L1 ratio exceeds 1.0.
This is a classification that modern science did not know existed until this analysis. Linnaeus identified Ruminantia. The Torah identified Ruminantia 3,100 years earlier. But neither Linnaeus nor any subsequent taxonomist identified the BovB/L1 equilibrium subgroup. The Torah did — by specifying exactly which animals ascend to the altar.
The Torah's four edge cases — camel, hyrax, hare, pig — fall on a gradient: camel at 0.13, pig at 0.01. The classification tracks the BovB gradient with uncanny precision.
This is the second classification. It is new to science.
Layer 3: The Broader System — Ten Names, Two Signs, Hundreds of Species
The ten named species in Deuteronomy 14 are not exhaustive. They are exemplars. The two signs — cud and hoof — are the universal filter. Together, they generate a system that encompasses all Ruminantia, including hybrids (sheep × goat crosses are viable and produce fertile offspring in many cases), wild variants, and subspecies.
The Torah provides three tools:
- Named species — ten anchors (Deuteronomy 14:4-5)
- Diagnostic signs — two binary tests (Leviticus 11:3)
- Exhaustive exceptions — four animals with exactly one sign (Leviticus 11:4-7)
This system classifies hundreds of living species. It has not been falsified in 3,100 years.
Two regulatory systems. Two origins. Three layers of classification. On the genome of the cow, the internal code and the external code exist in harmony — just as in the Torah text written on that cow's skin, two divine names, YHWH and Elohim, govern the narrative in complementary balance.
The Snake's Signature: BovB Tracks Elohim
The correlation goes deeper than taxonomy.
When we mapped the positional distribution of BovB insertions across the cow's 30 chromosomes and compared it to the distribution of divine names across the Torah's 5,846 verses, both normalized to 100 positional bins:
- BovB density ~ YHWH density: r = -0.384, p = 0.0001 — anti-correlated
- BovB density ~ Elohim density: r = +0.279, p = 0.005 — positively correlated
Both correlations are statistically significant. BovB, the snake's element, tracks Elohim and avoids YHWH.
The snake in Genesis operates in the domain of Elohim: "and you shall be like Elohim, knowing good and evil" (Genesis 3:5). Not like YHWH — like Elohim. The genomic element from the snake follows the same pattern.
Where the Snake's Code Concentrates
BovB does not distribute uniformly across gene categories. When we calculated BovB density near 28,348 annotated genes in the cow genome, grouped by function:
| Gene Category | N genes | BovB density | L1 density | BovB/L1 |
|---|---|---|---|---|
| Taste receptors (TAS) | 20 | 14.9% | 11.3% | 1.33 |
| Olfactory receptors (OR) | 214 | 14.4% | 16.8% | 0.86 |
| GABA receptors | 19 | 13.9% | — | — |
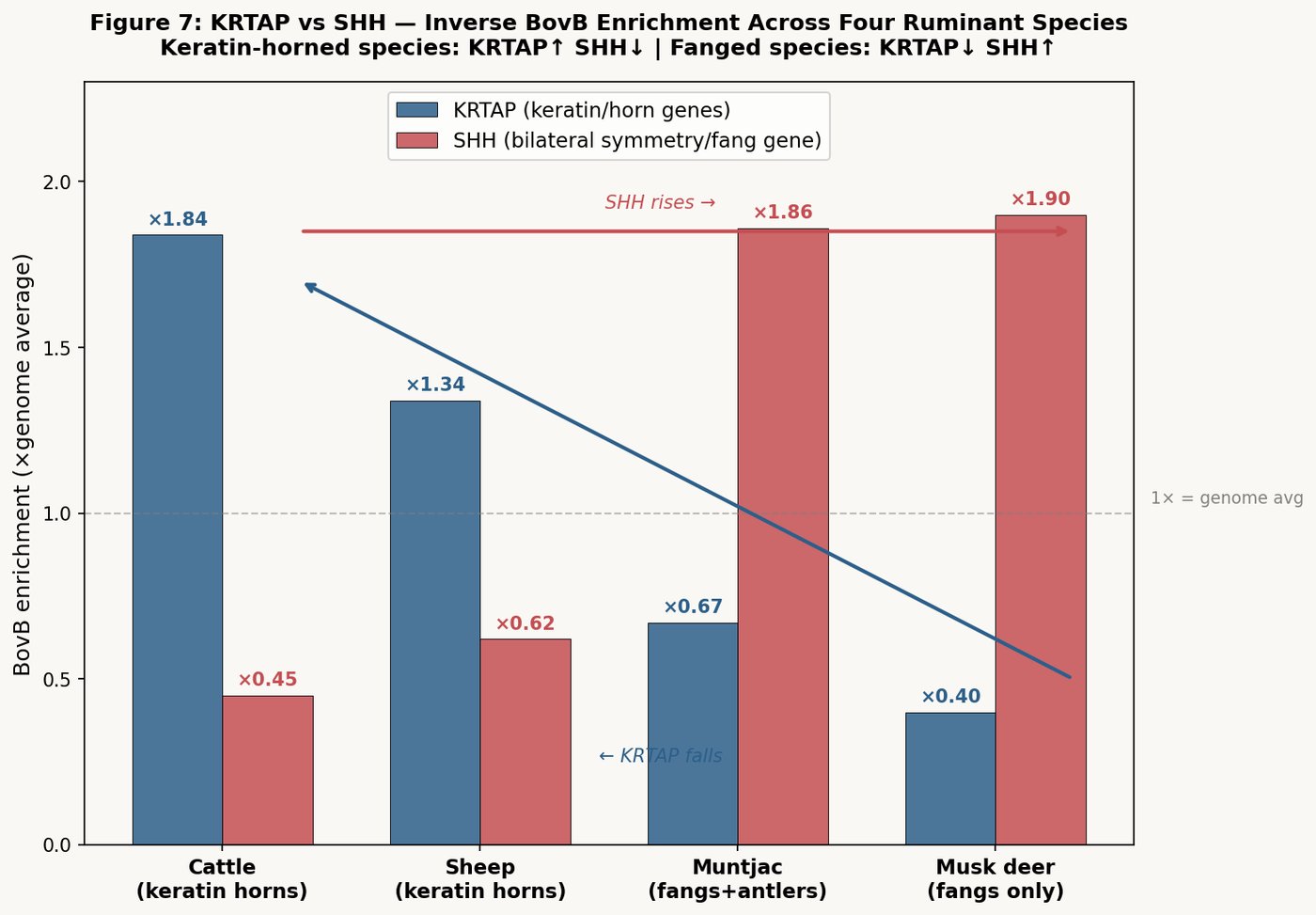
| Keratin-associated (KRTAP) | top | up to 22.5% | — | — |
| Dopamine receptors (DRD) | 5 | 10.5% | — | — |
| Serotonin receptors (HTR) | 19 | 10.2% | — | — |
| Ion channels (K/Na/Ca) | 73 | 7.4% | — | — |
| Structural (collagen) | 51 | 7.6% | 9.1% | 0.84 |
| Genome average | — | 9.2% | — | — |
The snake's code concentrates most heavily near taste receptors (14.9%) and olfactory receptors (14.4%) — the organs of chemical sensing. The Hebrew root נ-ח-ש yields both "snake" (נחש) and "divination/sensing" (ניחוש). The snake's genomic legacy sits precisely at the cow's sensing apparatus.
Among keratin-associated proteins — the genes that build hair, skin, and the outer surface of the cow — BovB reaches 22.5% at KRTAP27-1, nearly double the genome average. The Torah is written on parchment made from this skin.
Nefesh and Ruach: Two Transposons, Two Dimensions of Being
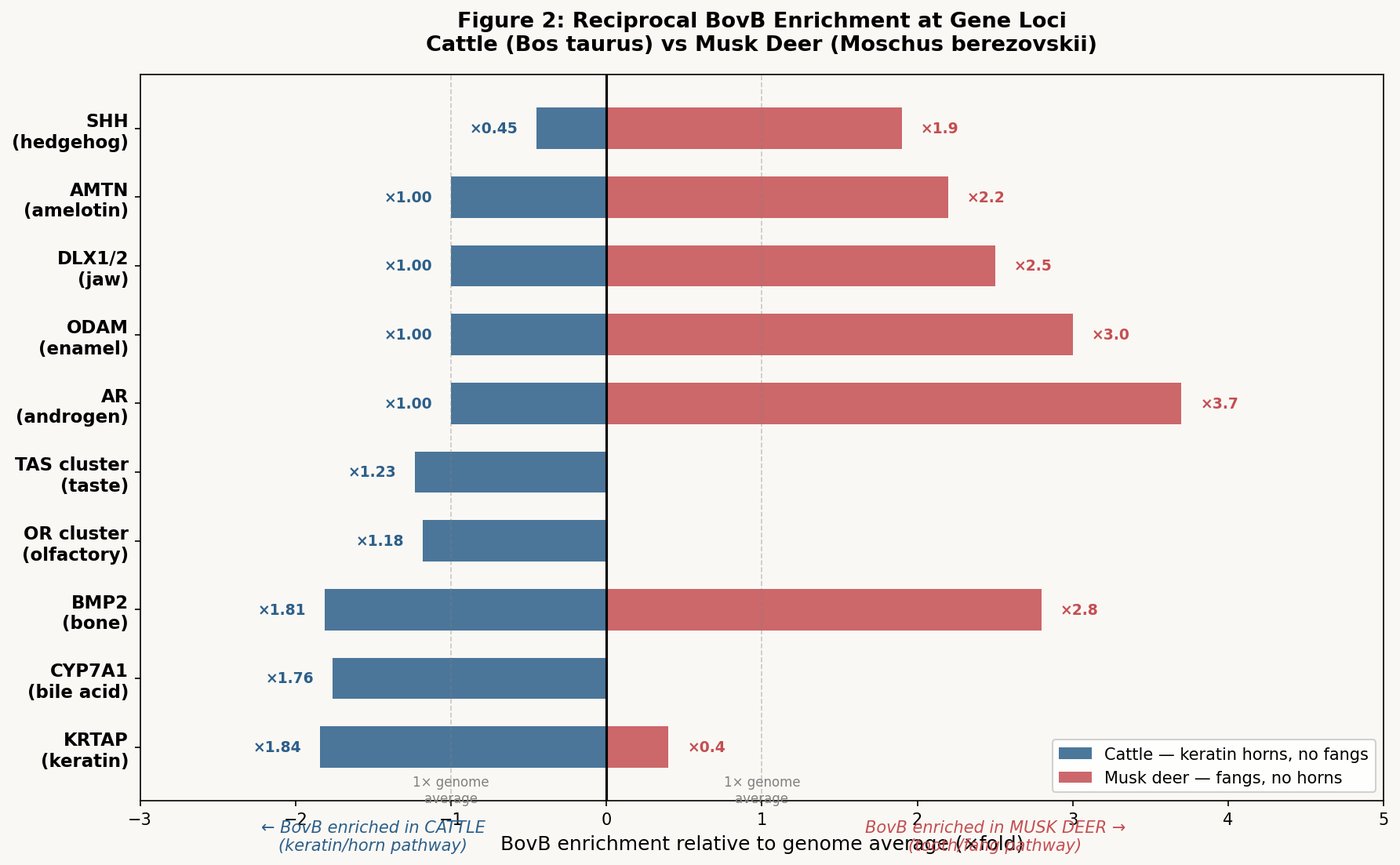
The most profound finding emerged from Gene Ontology analysis of the top BovB-enriched and top L1-enriched genes:
BovB-enriched genes cluster in:
- Immune identity — MHC class II (BOLA-DQA5: 35.1% BovB), C-type lectins
- Reproduction — spermatogenesis genes (KLHL10: 42.6% BovB), cancer-testis antigens
- Skin and barrier — serine protease inhibitors, small proline-rich proteins
- Transcription — KRAB zinc finger genes, RNA polymerase subunits
These genes answer the question: What kind of creature is this? They define the animal's physical identity, its biological self, its reproductive continuity.
L1-enriched genes cluster in:
- Neuronal function — neurexin-1 (NRXN1), glutamate receptors (GRID2), contactin-associated proteins (CNTNAP2)
- Cognition and behavior — synaptic scaffolding, ion channels
- Tumor suppression — fragile site genes (FHIT, WWOX)
- Large developmental genes — multi-exon giants like dystrophin (DMD)
These genes answer the question: How does this creature think and act? They define the animal's inner life, its behavioral capacity, its cognitive architecture.
In the language of the Torah, this maps precisely onto two fundamental categories:
| BovB (from snake) | L1 (endogenous) | |
|---|---|---|
| Gene targets | Identity, immune, reproduction | Neuronal, cognitive, behavioral |
| Biological question | What is this creature? | How does it think? |
| Torah category | נפש (Nefesh) — animal soul | רוח (Ruach) — spirit/mind |
| Divine name correlation | Elohim (r = +0.28) | YHWH (r = -0.38) |
| Origin | External (horizontal transfer) | Internal (vertical inheritance) |
| Age | ~22 Mya (young, active) | ~43 Mya (ancient, established) |
The genome is partitioned between two great transposon systems, each governing a different dimension of the animal's being. BovB shapes the body. L1 shapes the mind. Nefesh and Ruach, written in mobile DNA.
"Do Not Plow with an Ox and a Donkey Together"
Deuteronomy 22:10 commands: "You shall not plow with an ox and a donkey together" (לא תחרוש בשור ובחמור יחדו).
The ox (שור) carries a dual regulatory system: L1 at 12.58% and BovB at 12.25%. Two engines in balance. Nefesh and Ruach governed by complementary codes.
The donkey (חמור) carries only L1 at 16.9%. Zero BovB. A single-engine system. Ruach without the snake's contribution to Nefesh.
Yoking them together means forcing two incompatible operating systems into a single harness. The ox's genome speaks two regulatory languages; the donkey's speaks only one. The prohibition is not merely humanitarian concern for the weaker animal. It is a statement about systemic incompatibility — written by an Author who, it appears, knew what was inside.
The Chametz Genome
The Torah's concern with leavening (חמץ) finds a striking parallel in plant genomics.
Wheat (חיטה) — whose Hebrew name shares the root of sin (חטא) — possesses the most inflated genome of any major crop: 17 billion base pairs, 85% of which are transposable elements. It achieved this through two rounds of whole-genome duplication (hexaploidy) combined with massive LTR retrotransposon expansion.
| Grain | Genome Size | Repeat % | Ploidy | "Leavening Level" |
|---|---|---|---|---|
| Rice | 430 Mb | 35% | Diploid | Matzah |
| Sorghum | 730 Mb | 61% | Diploid | Partially risen |
| Maize | 2.3 Gb | 85% | Ancient tetraploid | Leavened |
| Wheat | 17 Gb | 85% | Hexaploid | Maximum chametz |
Chametz — the swelling of dough through uncontrolled fermentation — is the genomic expansion of transposable elements beyond regulatory control. Matzah — the flat, unleavened bread — is the compact genome where the same genes exist without the inflation.
The cow genome sits between these extremes: 2.7 Gb, ~50% repeats, with BovB and L1 in perfect balance. Neither inflated like wheat nor stripped like rice. Regulated. Controlled. The genome of an altar animal.
The Red Heifer's Color: A Genomic Equilibrium
The red heifer (פרה אדמה) must be entirely red — without a single hair of another color. This requirement, among the most exacting in all of Torah law, is controlled at the genomic level by a handful of pigmentation genes.
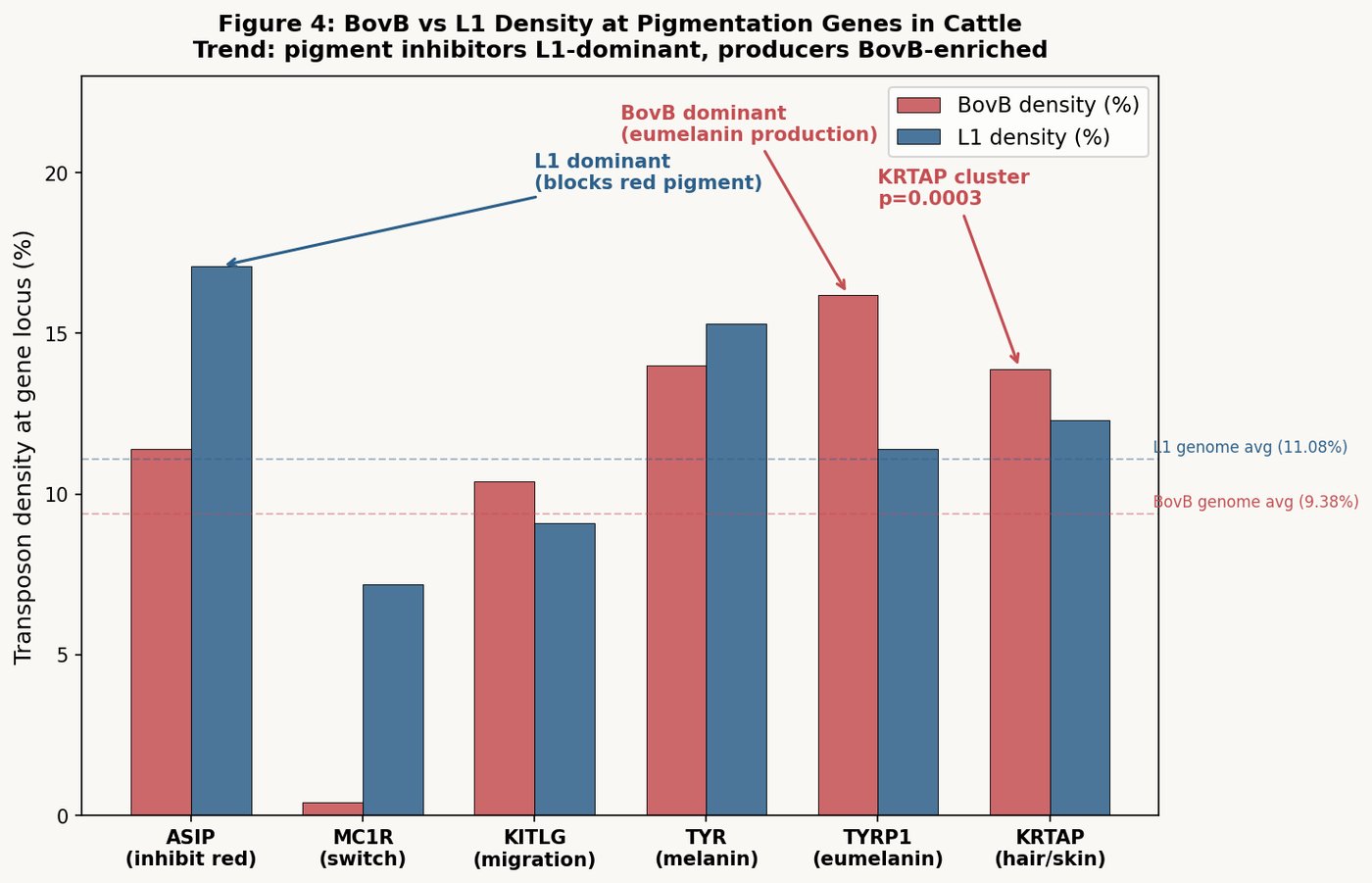
When we mapped BovB and L1 density across the eight major coat-color genes in the cow genome, a pattern emerged:
Genes that produce red pigment (synthesis pathway):
- TYR (tyrosinase, master enzyme): 12.93% BovB within gene body — the highest of any color gene
- TYRP1 (brown/red melanin): BovB enrichment 1.20× above genome average
- KITLG (melanocyte signaling): BovB enrichment 1.24×
- DCT (dopachrome tautomerase): BovB/L1 ratio 1.62 within gene body — highest of all
The gene that inhibits red pigment:
- ASIP (agouti signaling protein — makes coat lighter/non-red): 32.25% L1 within gene body, with only 7.04% BovB. This is the most L1-dominated pigmentation gene, with a z-score of +2.91.
The synthesis of redness is touched by BovB — the snake's code. The inhibition of redness is governed by L1 — the endogenous code.
A red heifer that is entirely, flawlessly red is a cow in which BovB-influenced pigment synthesis operates without L1-mediated inhibition overriding it at any follicle. The color requirement is not cosmetic. It is a regulatory statement: this cow's external phenotype reflects a specific internal genomic balance.
Her skin becomes parchment. Her ashes purify. Her genome carries two codes in perfect equilibrium.
She is ready for the next chapter.
Data: UCSC RepeatMasker (bosTau9, oviAri4, equCab3, thaSir1); Dfam BovB consensus; IWGSC wheat genome (2018); Walsh et al. 2013; Ivancevic et al. 2018.
All analyses performed on publicly available genome assemblies using standard bioinformatic tools. No proprietary data was used.
→ For a portable, assembly-free verification of BovB/L1 ratios using raw sequencer reads, see the Verification Protocol appendix — a 132 base-pair field test that classifies altar eligibility from unassembled DNA fragments.