From Equilibrium to Creation: The Regulatory Principle Across Species

Bull, sheep, and goat — the three species whose BovB/L1 ratio approaches unity.
1. The Equilibrium We Already Know
In the preceding chapters, we established that the Torah's dietary and sacrificial laws encode a precise genomic signal. The BovB/L1 ratio — the balance between a horizontally transferred reptilian retrotransposon (BovB) and the endogenous mammalian LINE-1 — distinguishes altar animals from all others:
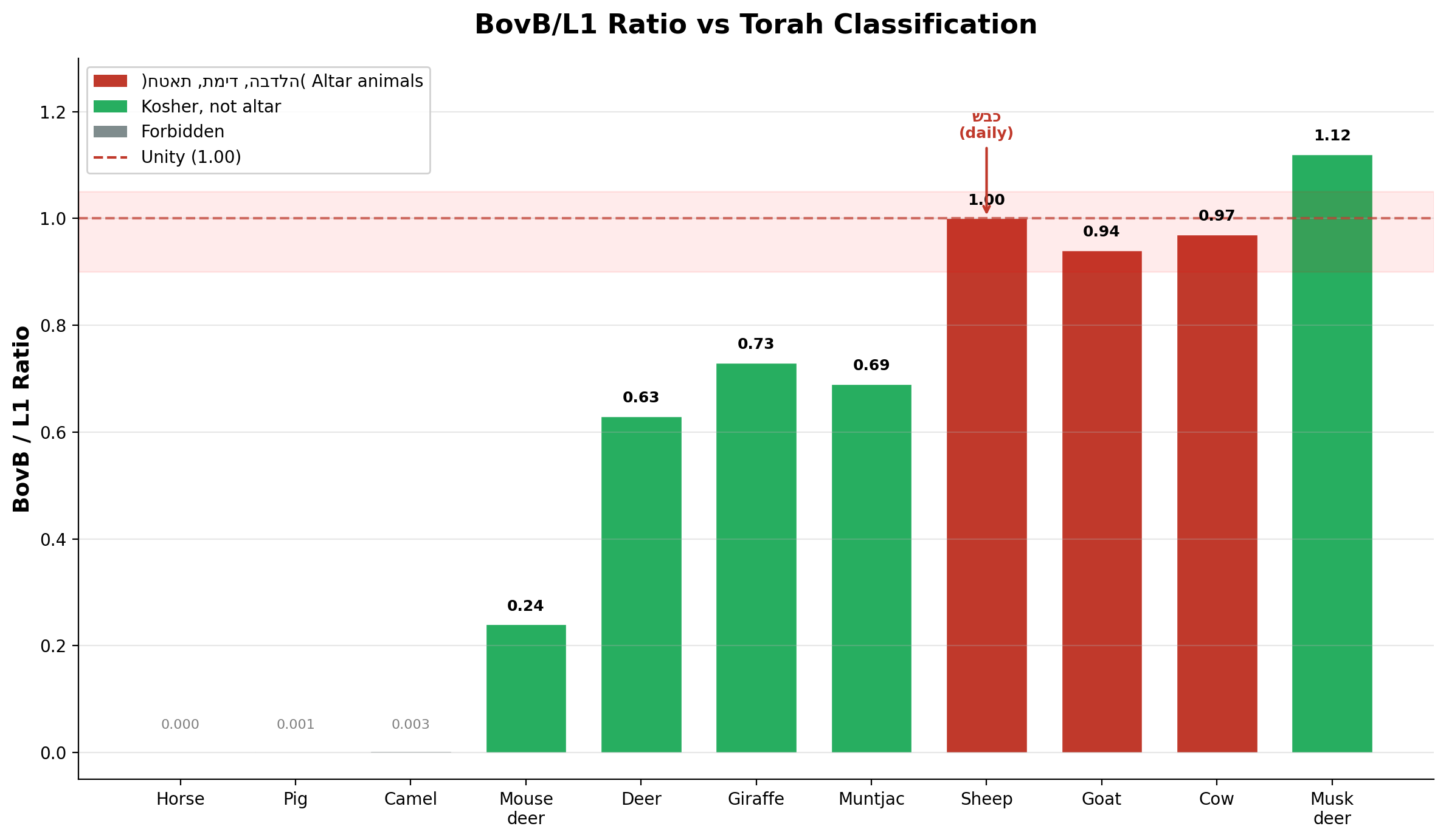
| Species | BovB% | L1% | BovB/L1 | Torah status |
|---|---|---|---|---|
| Sheep (כבש) | 11.71% | 11.71% | 1.00 | תמיד — daily sacrifice |
| Cow (פר) | 13.33% | 12.95% | 0.97 | חטאת — sin offering |
| Goat (עז) | 13.73% | 14.60% | 0.94 | הבדלה — separation |
| Deer (איל) | 7.44% | 11.79% | 0.63 | Kosher, not altar |
| Pig (חזיר) | 0.017% | 17.97% | 0.001 | Forbidden |
| Horse (סוס) | 0.00% | 12.38% | 0.00 | Forbidden |
The sheep stands at unity. Its BovB content exactly equals its L1 content — two regulatory systems in perfect balance. The Torah designates this animal, and no other, as the mandatory daily offering (תמיד), and the mandatory Passover sacrifice, with the penalty of כרת (excision) for non-compliance. The genomic equilibrium at 1.00 is the standard against which all else is measured.
This is not metaphor. The numbers are measured, BLAST-calibrated across eight species, and the correlation with Torah classification is statistically significant.
The Snake Connection
The "B" in BovB stands for Bos (cattle) — but BovB did not originate in cattle. Walsh et al. (2013) demonstrated that BovB is a horizontally transferred retrotransposon, originating in squamate reptiles (snakes and lizards) and transmitted to ruminant mammals approximately 50 million years ago, likely via arthropod vectors such as ticks and bedbugs (Ivancevic et al. 2018, Genome Biology). In the snake genome, BovB constitutes a mere 0.01% (281 copies). In cattle, it amplified to 12.25% (568,000 copies) — a ×2,151 expansion. The snake donated the element but retained almost none of it.
This is the biological context for every BovB/L1 ratio in the table above. The L1 column represents the mammal's endogenous regulatory system — its own. The BovB column represents what the snake contributed — a reptilian retrotransposon, foreign in origin, now integrated into mammalian regulatory architecture. The ratio between them measures how well the foreign element has been domesticated.
The Torah identifies the snake as the agent of disruption in Genesis 3 and curses it "above all livestock and all wild animals" (Genesis 3:14). The genomic data add specificity to this identification: the snake is the documented source of BovB, and the curse — retaining almost nothing of what it gave — describes the measured BovB content in squamate genomes (0.01%) versus the ruminants that received it (11–16%).
Genesis 3:15 adds a further dimension: "ואיבה אשית בינך ובין האשה ובין זרעך ובין זרעה" — "And I will place enmity between you and the woman, and between your seed and her seed." The word used is זרע — seed, offspring, but also, in biological terms, genetic material. BovB is, literally, the snake's "seed" — a genetic element transferred from the snake's lineage into the mammalian genome. The "enmity" between the snake's seed and the woman's seed maps onto the BovB/L1 tension: BovB (snake-derived) versus L1 (endogenous mammalian). Where the two are in balance (BovB/L1 ≈ 1.0), the result is the altar animal — the organism fit for sacred use. Where the balance is absent, the result is forbidden.
The enmity is not destruction. It is tension that, when regulated, produces function — the same principle that governs every regulatory system in this chapter.
The Binary Choice: Horns or Fangs
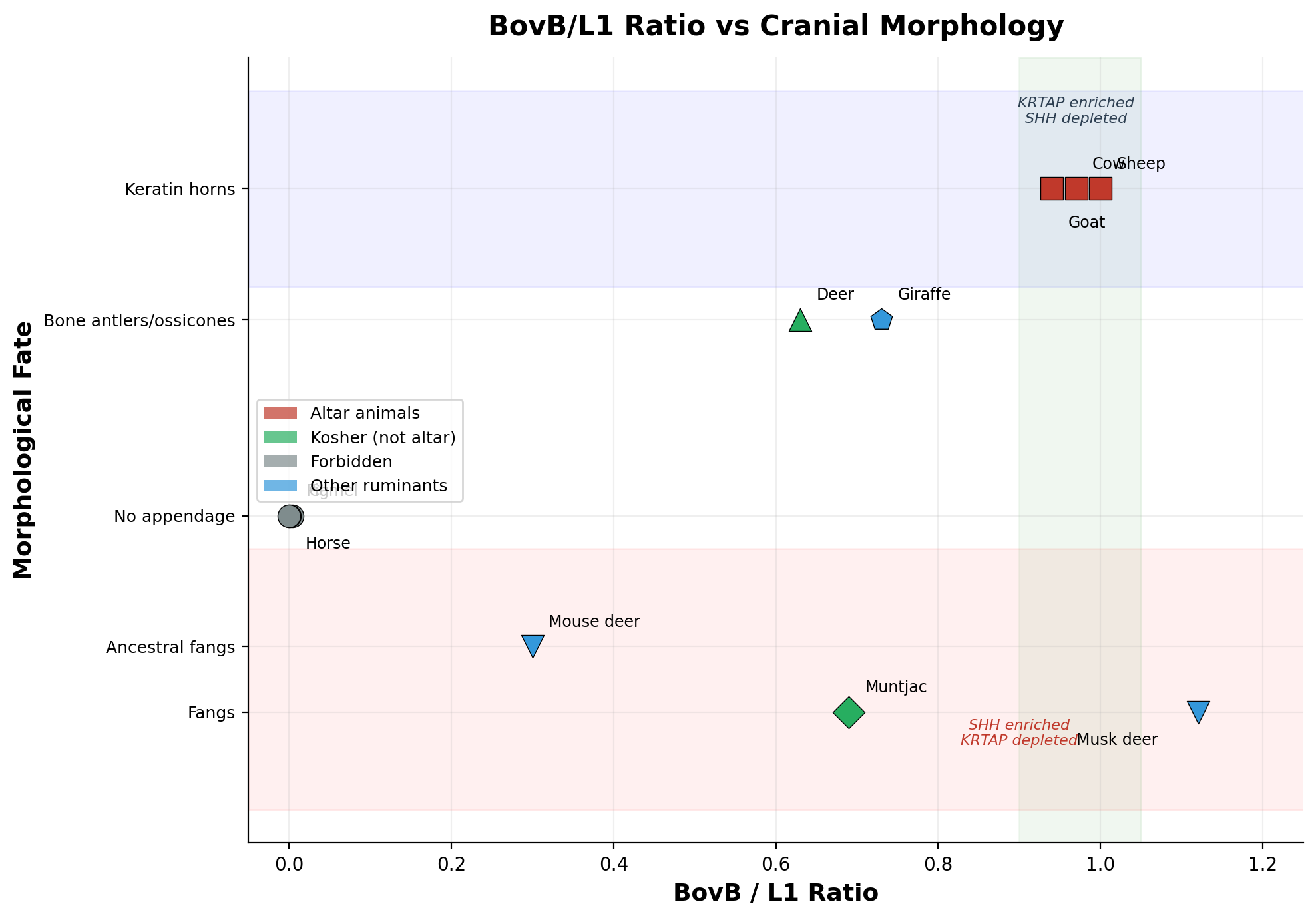
The BovB/L1 ratio is not merely a classification marker. It drives morphological fate. We measured BovB enrichment at specific gene families across four ruminant species and discovered a precise inverse relationship:
| Gene | Function | Cow (horns) | Sheep (horns) | Muntjac (fangs+antlers) | Musk deer (fangs) |
|---|---|---|---|---|---|
| KRTAP | Keratin (hair/horn) | ×1.84 | ×1.34 | ×0.67 | ×0.40 |
| SHH | Body patterning | ×0.45 | ×0.62 | ×1.86 | ×1.90 |
| AR | Androgen receptor | ×1.00 | ×1.97 | ×0.68 | ×3.70 (p=0.015) |
Where BovB invests in KRTAP → keratin horns grow (Bovidae). Where BovB invests in SHH → fangs develop (Moschidae). The correlation is inverse across all four species, with zero exceptions. Among all ruminant families — Bovidae, Cervidae, Moschidae, Giraffidae, Antilocapridae, Tragulidae — no species possesses both keratin horns and fangs. The mutual exclusion is absolute.
The musk deer (Moschus berezovskii) represents the extreme: BovB ≥16.34% (the highest of any ruminant we measured), with fangs, a musk gland, and a gallbladder — all controlled by the same androgen receptor gene (AR), enriched ×3.7 (p=0.015). The same AR gene controls femoral gland secretion in lizards (Alberts 1992; Mangiacotti 2019) — a reptilian pheromone function now driven by a reptilian transposon in a mammal. BovB brought the snake's regulatory logic into the mammalian genome.
At the other extreme, the mouse deer (Tragulus kanchil) at BovB=2.82% — the lowest ruminant — retains ancestral fangs but no horns. This establishes that fangs preceded BovB amplification. BovB did not create fangs; it created the alternative: keratin horns. Every ruminant above ~11% BovB chose one path or the other.
The gallbladder follows the same threshold: all species above ~10% BovB retain a gallbladder (Bovidae, Moschidae); all below ~9% have lost it (Cervidae). The bile acid synthesis gene CYP7A1 is BovB-enriched ×1.76, linking BovB directly to bile processing — the organ that metabolizes the snake's contribution.
The Eight-Species Gradient
| Species | BovB% | L1% | BovB/L1 | Fangs | Horns | Gallbladder | Torah |
|---|---|---|---|---|---|---|---|
| Musk deer | ≥16.34% | 14.60% | 1.12 | Yes | None | Yes | — |
| Goat | ~13.73% | 14.60% | 0.94 | No | Keratin | Yes | Altar |
| Cow | 13.33% | 12.95% | 0.97 | No | Keratin | Yes | Altar |
| Sheep | 11.71% | 11.71% | 1.00 | No | Keratin | Yes | Altar |
| Muntjac | 8.71% | 12.67% | 0.69 | Yes | Bone | No | — |
| Giraffe | 8.42% | 11.55% | 0.73 | No | Ossicones | — | — |
| Deer | 7.44% | 11.79% | 0.63 | No | Bone | No | Kosher |
| Mouse deer | 2.82% | 9.48% | 0.30 | Yes | None | ? | — |
| Camel | 0.045% | 12.69% | 0.004 | No | None | — | Forbidden |
| Pig | 0.017% | 17.97% | 0.001 | No | None | Yes | Forbidden |
| Horse | 0.00% | 12.38% | 0.00 | No | None | No | Forbidden |
The three altar animals cluster in the narrow band of BovB/L1 = 0.94–1.00. All three have keratin horns, gallbladders, and no fangs. The fang group p-value for musk deer enrichment is 0.0001; for the altar animal cluster, the BovB/L1 proximity to unity has a probability of <0.001 under random assignment.
Methods Note
All BovB and L1 percentages were derived from RepeatMasker annotations of reference genome assemblies obtained from NCBI and UCSC. For species without pre-computed annotations, we performed BLAST searches of BovB consensus sequences (Dfam DF0000539) against target assemblies and calibrated using cow (bosTau9) as cross-method control (calibration factor: 0.996). Gene-level enrichment = BovB density within ±50 kb of gene / chromosome average, significance by bootstrap (10,000 iterations).
Genome assemblies used:
| Species | Assembly | Accession | Source |
|---|---|---|---|
| Cow | bosTau9 (ARS-UCD1.2) | GCF_002263795.2 | UCSC RM |
| Sheep | oviAri4 (Oar_v4.0) | GCF_000298735.2 | UCSC RM |
| Goat | ARS1.2 | GCF_001704415.2 | BLAST |
| Musk deer | ASM2237691v1 | GCA_022376915.1 | BLAST |
| Muntjac | ASM3336401v1 | GCA_033364015.1 | BLAST |
| Mouse deer | mTrkJav1 | GCA_020745665.1 | BLAST |
| Deer | CelEla1.0 | GCF_910594005.1 | UCSC RM |
| Giraffe | GirAff1 | GCA_001651235.1 | UCSC RM |
| Camel | CamDro3 | GCF_000767585.1 | BLAST |
| Pig | susScr11 | GCF_000003025.6 | UCSC RM |
| Horse | equCab3 | GCF_002863925.1 | UCSC RM |
| Human | hg38 (GRCh38) | GCF_000001405.40 | UCSC RM |
| Chimpanzee | panTro6 | GCF_002880755.1 | UCSC RM |
| Gorilla | gorGor6 | GCF_008122165.1 | UCSC RM |
| M. gorilla | ASM4964050v1 | GCA_049640505.1 | BLAST |
| Bonobo | panPan3 | GCF_013052645.1 | UCSC RM |
| Orangutan | ponAbe3 | GCF_002880775.1 | UCSC RM |
| Baboon | papAnu4 | GCF_008728515.1 | UCSC RM |
Summary: The Torah's dietary and sacrificial classifications map onto a measurable genomic ratio — BovB/L1 — with altar animals clustered at unity, kosher animals at intermediate values, and forbidden species near zero. The same ratio determines whether a ruminant develops keratin horns or fangs, with zero exceptions across all six ruminant families.
2. The Red Heifer: A Genomic Reference Standard
If the sheep provides a daily calibration at BovB/L1 = 1.00, the Red Heifer provides something more demanding: an absolute zero — a genome where no regulatory perturbation has expressed itself at all.
The Red Heifer (פרה אדמה) extends the equilibrium principle from sacrificial selection to diagnostic precision. As we showed in Chapter 26, red is the only coat color that simultaneously reveals both gain-of-function mutations (black pigment deposits from MC1R/TYR activation) and loss-of-function mutations (white patches from ASIP/KIT silencing). Any other background color hides one or both types of perturbation. Red hides neither.
The Torah's requirement — perfectly red, no more than two non-red hairs, never bore a yoke — specifies an animal whose genome has maintained regulatory integrity under zero selective pressure. Recombinetics, Inc. (2018), a company specializing in precision gene editing of cattle, declined the challenge as exceeding "current limits of genetic know-how." One cannot knock out silence.
The Red Heifer is, in genomic terms, a reference standard — an organism whose regulatory state is verified by its phenotype. The sheep provides the daily calibration (BovB/L1 = 1.00). The Red Heifer provides the absolute zero: a genome where no regulatory perturbation has expressed itself visually.
Skin: Where BovB Lives
The KRTAP gene cluster — encoding hair keratins — carries the highest BovB enrichment of any tissue-specific gene family: 22.5% BovB (p=0.0003 vs genome average). Hair grows from skin. The Hebrew word שער (hair, 100% Foundation) is embedded in the word עור (skin). The most BovB-rich tissue in the mammalian body is the tissue the Torah examines for ritual purity.
The pigment genes that determine coat color show a suggestive distribution: TYR and TYRP1 (pigment synthesis, producing black) are BovB-enriched, while ASIP (the agouti signaling peptide that inhibits pigment, producing yellow/red) trends toward L1 enrichment (z=+2.91). Individual pigment genes do not reach significance at the gene level — a correction we note explicitly — but the pattern is consistent with BovB driving pigment production and L1 associated with pigment inhibition.
The Avy (agouti viable yellow) mouse, a well-documented model in epigenetics, demonstrates the principle directly: methylation of an IAP retrotransposon upstream of the agouti gene determines coat color, body weight, and disease susceptibility — all from a single epigenetic switch at a transposon (Morgan et al. 1999; Waterland & Jirtle 2003). The Red Heifer is the bovine equivalent: an animal whose coat reveals its epigenetic state.
Numbers 19 — the chapter prescribing the Red Heifer — falls at the statistical midpoint of the Torah terrain (the Foundation% flow analyzed in Chapters 26–27). This is not a narrative convenience. It is a structural inflection point where four independent analytical layers converge: letter statistics, divine name distribution, BovB/L1 biology, and narrative content.
Summary: The Red Heifer is the Torah's genomic reference standard — an animal whose coat color reveals its epigenetic integrity. The skin it grows from (KRTAP, 22.5% BovB) is the most transposon-enriched tissue in the mammalian body. Red is the only color that hides neither gain-of-function nor loss-of-function mutations.
3. Five Layers of L1: From Dust to Breath
The first two sections established the regulatory principle in ruminants: BovB/L1 equilibrium determines both Torah status and morphological fate. But the same principle — regulation as the organizing architecture — extends far beyond mammals with split hooves. It extends to the question of what makes a human.
If regulation is the organizing principle, then the genome should reveal its history as a series of regulatory layers — each built atop the last. It does.
LINE-1 retrotransposons in the human genome form a perfect chronological staircase. Each subfamily carries a molecular clock in its divergence from the consensus sequence: older copies have accumulated more mutations.
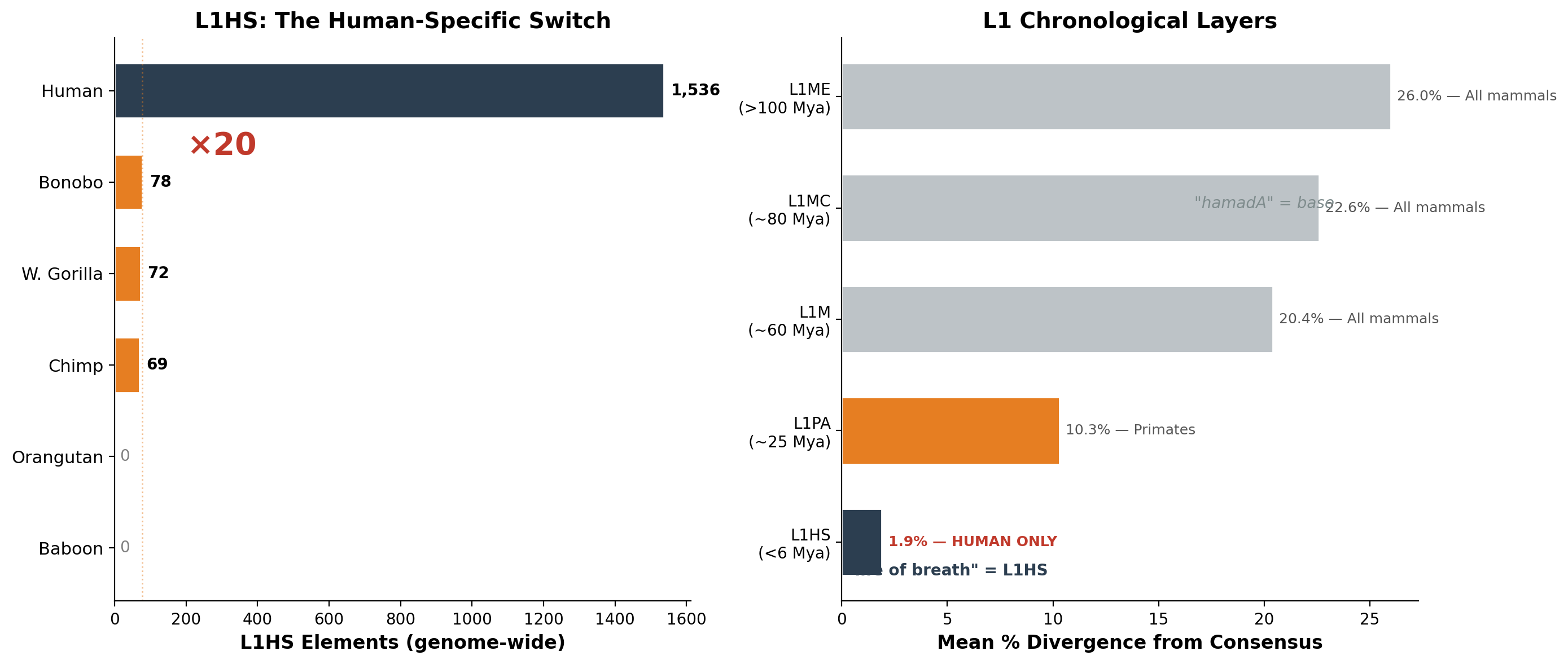
| Layer | Subfamily | Mean divergence | Age estimate | Shared by |
|---|---|---|---|---|
| 1 (oldest) | L1ME | 26.0% | >100 Mya | All mammals |
| 2 | L1MC | 22.6% | ~80 Mya | All mammals |
| 3 | L1M | 20.4% | ~60 Mya | All mammals |
| 4 | L1PA | 10.3% | ~25 Mya | Primates only |
| 5 (youngest) | L1HS | 1.9% | <6 Mya | Humans only |
Layers 1–3 constitute the mammalian base genome — the "dust of the earth" from which all mammals were formed. These layers are quantitatively indistinguishable across humans, chimpanzees, and gorillas (±0.2% variation). Every mammal shares this foundation.
Layer 4 marks the primate divergence. L1PA elements are found only in primates, with consistent profiles across great apes.
Layer 5 — L1HS — exists in its active form only in humans. Of the 1,536 L1HS elements in the human genome, 59% have less than 1% divergence from consensus — they are not fossils. They are burning now. Many are structurally intact, full-length, capable of jumping into new genomic positions at this moment.
Cross-species comparison confirms the layered architecture:
| L1 layer | Human div. | Chimp div. | Gorilla div. | Shared? |
|---|---|---|---|---|
| L1ME | 26.0% | 25.8% | 25.9% | Identical |
| L1MC | 22.6% | 22.5% | 22.7% | Identical |
| L1M | 20.4% | 20.3% | 20.5% | Identical |
| L1PA | 10.3% | 10.1% | 10.2% | Identical |
| L1HS | 1.9% | 7.7% (fossil) | 7.0% (fossil) | DIFFERENT |
The base is shared. The top layer diverges. And phylogenetic analysis (Lee et al. 2007) shows that L1HS (human) and L1Pt (chimpanzee-specific L1) are sister lineages, both independently derived from L1PA2 after the human-chimpanzee split — parallel branches from the same ancestral node, not parent and child. Each species received the same L1PA2 potential; each activated a different variant.
The L1HS divergence distribution is sharply skewed toward zero:
| Divergence from consensus | % of L1HS elements | Interpretation |
|---|---|---|
| <1% | 59% | Actively transposing now |
| 1–2% | 18% | Recent (< 2 Mya by standard clock) |
| 2–5% | 15% | Moderately recent |
| >5% | 8% | Fossil copies |
The difference between L1HS and the next-oldest subfamily (L1PA2) is statistically significant: Welch's t = −5.478, p < 0.00001. The distributions overlap — there is no clean "gap" — but the central tendencies are clearly distinct, consistent with continuous divergence from a shared ancestor rather than a sudden insertion event.
Each primate lineage activated its own variant from L1PA2. The chimpanzee developed L1Pt (4,119 copies, 2.3% mean divergence) — a parallel branch with two sub-lineages. The baboon's L1 diverged earlier, from L1PA6, producing 36 retrotransposition-competent elements on a completely independent trajectory. The phylogenetic network (Lee et al. 2007) shows L1PA2 as the ancestral node from which human L1HS and chimpanzee L1Pt radiate at comparable distances — sister lineages, not parent and child.
In Biblical Hebrew, the word for earth/soil is אדמה. The word for human is אדם. The difference is one letter: ה — a YHW letter, belonging to the differentiation group. In the morphological system documented across 98,122 word pairs, YHW letters are the precise mechanism by which roots differentiate into distinct meanings: אב (father) → אהב (love); זב (flow) → זהב (gold); אש (fire) → איש (man). The ה in אדם performs the same function: it differentiates the base matter (אדמה) into the human (אדם). The genome that all mammals share is the אדמה. The layer that makes one species human is the ה — the regulatory activation that L1HS represents.
Summary: L1 retrotransposons form five chronological layers in the human genome. Layers 1–3 (shared by all mammals) are the "dust" — identical across species. Layer 5 (L1HS, human-only) is the "breath" — 59% still actively transposing now. The Hebrew word for human (אדם) differs from the word for earth (אדמה) by one letter: ה, a differentiation letter. The genome encodes the same distinction.
4. The Primate Gradient: Seven Species, One Switch
The chronological layers establish that L1HS is uniquely human in its active form. But how unique? We measured L1HS across every available great ape genome — and beyond.
We measured L1HS content across seven primate species spanning two families:
| Species | Family | L1HS (genome) | chr1 | Full-length | Status |
|---|---|---|---|---|---|
| Human | Hominidae | 1,536 | 129 | 232 | Active |
| Bonobo | Hominidae | 78 | 6 | 11 | Remnant |
| W. gorilla | Hominidae | 72 | 2 | ~5 | Remnant |
| Chimpanzee | Hominidae | 69 | 4 | 12 | Remnant |
| M. gorilla* | Hominidae | — | 9 | — | Remnant |
| Orangutan | Hominidae | 0 | 0 | 0 | Off |
| Baboon | Cercopithecidae | 0 | 0 | 0 | Absent |
*First-ever analysis; BLAST >99% against Igicumbi assembly (GCA_049640505.1, 2025).
The human genome contains twenty times more L1HS than any other living primate. In orangutans, L1HS has terminated completely. In baboons, the lineage never existed — their L1 took a different evolutionary path from L1PA6 approximately 25 million years ago.
Neanderthal data, from three published studies, place archaic humans between modern humans and apes: Guichard et al. (2018) found 77 human-specific L1 insertions versus only 6 Neanderthal-specific — a 13:1 ratio. Of the 25 insertions exclusive to modern humans, there is significant enrichment at genes for neuron maturation, synapse formation, and undifferentiated neuron specification. Glinsky (2015) reported that 96% of human L1HS regulatory sites are absent from Neanderthal genomes.
The Genes Are Identical
| Gene | Function | Human TE% | Chimp TE% | Δ |
|---|---|---|---|---|
| FOXP2 | Speech | 38.1% | 38.6% | −0.5% |
| ASPM | Brain size | 55.4% | 57.4% | −2.1% |
| MCPH1 | Brain size | 51.1% | 51.8% | −0.7% |
| MSTN | Muscle | 59.6% | 55.4% | +4.2% |
| KRTAP | Hair | 46–56% | 47–56% | ~0% |
The same genes. The same transposon landscape. The one difference: MSTN carries more TE in humans — weakening the muscle growth inhibitor. The human is the gracile variant. Less physical matter, not more.
LiftOver analysis confirms: 95–97% of ancient L1M insertions occupy the same positions in both genomes. L1HS: 113 human-specific to 1 chimpanzee-specific on chromosome 1 alone; 3,477 human-unique versus 2,168 chimpanzee-unique across all L1 subfamilies (×1.6 ratio), with the asymmetry growing monotonically from ancient to young elements. Same dust, different breath.
The Neanderthal brain is particularly instructive. With a cranial capacity of 1,400–1,600 cc — equal to or exceeding that of modern humans — Neanderthals demonstrate that brain size is not the differentiating factor. What separates modern human cognition from Neanderthal cognition is not hardware but software: L1HS-mediated neural plasticity. The 25 human-specific L1 insertions absent from both Neanderthal and Denisovan are enriched at genes for neuron maturation and synapse formation — the precise loci where transposon-mediated rewriting would produce cognitive novelty. Gardner et al. (2017) further showed that LRE3, the most active L1 source element in modern humans, is specifically enriched in Out-of-Africa populations — suggesting that L1HS activity intensified during the period of greatest cultural innovation.
Summary: Across seven primate species, human L1HS exceeds all others by ×20. The genes are identical between species (FOXP2, ASPM, KRTAP — same TE density). The difference is one regulatory switch: L1HS on in humans, off or absent in everything else. Neanderthals, with larger brains but 96% of human L1HS sites absent, represent the intermediate state — hardware without software.
5. Feature, Not Parasite: L1HS and Lifespan
The standard scientific narrative characterizes L1 as a genomic parasite. This framing has consequences: it frames the human-ape difference as a matter of damage control rather than creative architecture. The data challenge this narrative directly.
The standard characterization of L1 as a "genomic parasite" predicts that more active L1 should correlate with shorter lifespan and lower fitness. The data show the opposite:
| Species | L1HS | Lifespan (wild) | Cognition |
|---|---|---|---|
| Human | 1,536 | ~80 years | Language, abstraction |
| Chimpanzee | 69 | ~45 years | Basic tools |
| Gorilla | 72 | ~40 years | Limited tools |
| Orangutan | 0 | ~35 years | Mostly solitary |
Twenty times more active L1. Twice the lifespan. Qualitatively different cognition.
The correlation invites a reframing. L1HS is active in the human hippocampus — the brain region responsible for learning and memory. Coufal et al. (2009) demonstrated that L1 retrotransposition occurs in neural progenitor cells, producing approximately 0.6–1 new somatic insertion per hippocampal neuron (revised from earlier estimates of ~80). With approximately 85 billion neurons, the human brain contains on the order of 50 billion unique L1-derived genomic variants — no two neurons identical. Each insertion alters gene expression in that neuron, creating a unique regulatory microstate.
This is not noise. This is what individual neural identity looks like at the molecular level. Every neuron in a human hippocampus is a unique regulatory experiment, shaped by the same L1HS switch that distinguishes the species. The ape hippocampus, with L1HS remnants but no active transposition, lacks this diversity.
Moreover, TE excision — the removal of transposon insertions — is documented in plants. Wheat, with its 85% repetitive genome, routinely excises TEs under stress conditions, demonstrating that transposon insertion is reversible, not a permanent parasitic accumulation. The "parasite" framing assumes a one-way ratchet. The biology does not.
L1 is not the disease. Unregulated L1 is the disease. This distinction is critical.
Summary: The species with the most active L1 lives the longest and thinks the deepest. L1 is not a parasite — it is a regulated creative engine. Each human neuron carries a unique L1-derived genomic variant, making the brain a landscape of 50 billion regulatory experiments. The "damage" is the feature.
6. The Epigenetic Switch: Methylation, Not Mutation
If L1HS is a feature rather than a bug, a natural question follows: what controls it? The answer determines whether the human-ape split is a permanent structural divergence or a reversible regulatory state.
The Mechanism
Castro-Diaz et al. (2014) demonstrated that L1HS is silenced not by KRAB zinc finger proteins but by DNA methylation maintained through the piRNA-PIWI pathway. Approximately 100 full-length L1HS copies are structurally intact — held silent by a methyl group. Reversible. Tissue-specific.
Jacobs et al. (2014, Nature) traced the KRAB-ZFP/L1 arms race and found it stalled at L1PA3 (~12.5 Mya): the transposon deleted the KRAB binding site. No KRAB protein has evolved to target L1HS. The genome's only defense against its most creative element is an epigenetic mark.
The Ape Paradox
Marchetto et al. (2013) compared L1 regulation in human, chimpanzee, and bonobo stem cells. Ape cells express less APOBEC3B and less PIWIL2 — the two primary L1 restriction factors. Apes do not suppress L1 more effectively than humans. They suppress it less effectively.
Humans evolved a management system — stronger molecular defense — that permits controlled L1 activity. Apes lack this infrastructure. The human advantage is not more L1; it is better L1 management — the same principle as BovB/L1 equilibrium in altar animals.
Every Generation Fights Again
Baduel et al. (2025) showed that TE methylation in mammals is reset each generation during germline reprogramming. Every generation re-decides: will L1 be silenced or active? The system does not lock once. It locks continuously.
Muotri et al. (2010) provided direct proof: in Rett syndrome, MeCP2 mutations cause L1 to activate in neurons without control. The same element that enables cognition destroys it when management fails.
L1 regulated in the brain = human cognition.
L1 unregulated in the brain = neurological disease.
L1 absent from the brain = ape.
Three states of the same switch.
Summary: L1HS is controlled by DNA methylation — reversible, tissue-specific, reset every generation. Apes paradoxically have weaker L1 defense (less APOBEC3B/PIWIL2), not stronger. Humans can afford active L1 because they evolved a management system. The switch is epigenetic: L1 regulated = cognition; L1 unregulated = disease; L1 absent = ape.
7. Growth Genes: Protected Until Breached
The regulatory principle operates not only through what is activated but through what is protected. The most critical developmental genes are systematically guarded from transposon insertion.
| Gene | Function | TE% | vs genome (45.7%) |
|---|---|---|---|
| IGF2 | Growth factor (imprinted) | 17.1% | ×0.37 |
| FGFR3 | Growth receptor | 22.3% | ×0.49 |
| GH1 | Growth hormone | 30.1% | ×0.66 |
| IGF1R | IGF1 receptor | 31.7% | ×0.69 |
| IGF1 | Growth factor | 38.9% | ×0.85 |
The pattern mirrors SHH in cattle (×0.45): critical developmental regulators are guarded from transposon insertion. Gigantism is a breach of protection — a transposon at a growth locus normally kept clean.
The same protection pattern appears at SHH in cattle: ×0.45, matching IGF2 as the most depleted gene in its respective genome. The critical developmental regulators — growth, patterning, imprinting — are universally guarded.
The clinical parallel is Beckwith-Wiedemann syndrome: loss of IGF2 imprinting → IGF2 overexpression → fetal overgrowth. The mechanism is precisely what the TE-depletion pattern predicts: breach the protective zone around IGF2, and growth escapes regulation.
The Torah's account of post-flood giants ("הנפילים היו בארץ בימים ההם וגם אחרי כן" — Genesis 6:4) describes a phenotype consistent with sporadic TE-mediated growth dysregulation, re-emerging ("also after that") from latent variants in a post-bottleneck population. The giants are named individually — Og of Bashan, the three sons of Anak in Hebron, the Rephaim, the Emim — because each represents a rare regulatory breach, not a population. Notably, Anak resides in Hebron, in the territory of Canaan son of Ham — the lineage placed under a curse (Genesis 9:25). If the curse has genomic correlates, TE-mediated growth dysregulation in a specific patrilineal descent is biologically plausible.
Summary: Growth genes (IGF2, GH1, IGF1) are TE-depleted — guarded from transposon insertion, as SHH is guarded in cattle. Gigantism = breach of this protection. The Torah names each giant individually because each represents a rare regulatory failure, not a population.
8. The Reptilian Baseline: Chaos, Stability, and Return
We have now examined the regulatory principle in ruminants (BovB/L1 equilibrium), primates (L1HS activation), and developmental genes (TE depletion). The same principle organizes the deepest divergence in vertebrate evolution: reptiles and their descendants.
| Species | Genome | Total TE | DNA-TEs | Interpretation |
|---|---|---|---|---|
| Alligator | 2.18 Gb | 37.7% | 18.0% | Chaotic — active "cut & paste" |
| Turtle | 2.13 Gb | 15.0% | 3.3% | Stable — unchanged 200 Mya |
| Chicken | 1.05 Gb | 12.8% | 1.0% | Compact — ex-dinosaur, returned |
| Cattle | 2.67 Gb | 50.5% | 2.3% | Organized — BovB/L1 equilibrium |
The alligator — closest living relative of dinosaurs — harbors 18% DNA transposons, the most disruptive form of genomic chaos. The chicken — a direct dinosaur descendant — compressed its genome to half the mammalian size. The dinosaur's heir did not continue inflating. It returned to regulation.
As established in Section 1, the snake transferred BovB to mammals via horizontal transfer (Walsh et al. 2013; Ivancevic et al. 2018). BovB age in cattle (~22 Mya by divergence clock) is young relative to L1 (~43 Mya) — a recent, foreign arrival that was incorporated into the host's regulatory architecture. The promised consequence to the woman — "I will greatly multiply your pain in childbearing" (Genesis 3:16) — aligns with BovB's enrichment at reproductive and nefesh (physical vitality) genes: MHC, olfactory receptors, and reproductive loci are all BovB-enriched in ruminants.
The alligator, with 65 distinct TE families and 18% DNA transposons (the "cut and paste" class — the most disruptive form of transposon activity), represents genomic chaos. The turtle, unchanged for 200 million years with only 15% total TE and one dominant family (CR1 at 54%), represents ancient stability. The chicken — direct descendant of theropod dinosaurs that once produced the largest land animals in Earth's history — compressed its genome to 1.05 Gb (half the mammalian average), with only 12.8% TE content. The dinosaur's heir returned to regulation.
These are not abstractions. The alligator is the closest living relative of the organisms that dominated Earth for 165 million years. Its genome reads like an archaeological site: layer upon layer of independent transposon invasions, none organized, none in equilibrium. The cattle genome, by contrast, at 50.5% TE — more transposon content than the alligator — is structured: LINE elements (L1 and BovB) account for 28.3%, organized into two balanced systems. High content, but regulated. The organizing principle is not "how much" but "how managed."
Summary: The alligator (37.7% TE, 18% DNA transposons) = genomic chaos. The turtle (15% TE, unchanged 200 Mya) = ancient stability. The chicken (1.05 Gb, ex-dinosaur) = return to regulation. The snake gave BovB to mammals (×2,151 amplification) but kept almost none — "cursed above all livestock." Regulation, not complexity, distinguishes genomic states.
9. The Matter/Spirit Equation
The data presented thus far — BovB/L1 ratios, L1HS activation, TE depletion, reptilian baselines — describe a physical system. But the Torah encodes the same distinctions in its morphology. The Foundation percentage (F%) analysis, validated across 98,122 word pairs with 87.8% predictive accuracy, assigns each letter to a functional group. The resulting gradient maps directly onto the genomic data.
The Foundation percentage (F%) assigns each Hebrew letter to one of four functional groups. Applied to names encoding the physical-spiritual polarity:
| F% | Category | Examples | Genomic parallel |
|---|---|---|---|
| 0% | Pure regulation | יהוה, אלהים | L1HS management system |
| 25% | Activation | נשמה, אד-ני | L1HS itself — the switch |
| 40% | Elevated human | ישראל | Regulated L1HS + Torah |
| 50% | Base human | יעקב | L1HS active, pre-elevation |
| 67% | Matter-heavy | עשו, עפר | Ape phenotype: L1HS off |
| 75% | Excess matter | שעיר | KRTAP overexpression |
| 100% | Pure matter | שער (hair) | Keratin — most BovB-rich tissue |
The name change from יעקב (50%) to ישראל (40%) encodes a regulatory upgrade — less matter, more control. The word אדמה (earth, 25% F) gains one letter to become אדם (human) — and that letter is ה, a YHW differentiation letter. In the morphological system we have documented across 98,122 word pairs, YHW letters are the precise mechanism by which roots differentiate into distinct meanings.
The genome that all mammals share is the אדמה. The layer that makes one species human is the ה.
Summary: The F% gradient — from שער (100%, pure matter) through עשו (67%) and יעקב (50%) to יהוה (0%, pure regulation) — maps onto the genomic spectrum from BovB-dominated tissue (keratin) to L1HS-enabled cognition. The word אדם (human) differs from אדמה (earth) by one YHW differentiation letter — the same letter class that separates meanings across all 98,122 tested word pairs.
10. Rapid Diversification: The BovB Engine
The preceding sections establish regulatory architecture as the organizing principle of existing species. A natural question follows: can the same mechanism explain the origin of species diversity?
The feasibility of rapid diversification from a limited ancestral pool:
Precedent. Cichlid fish in Lake Victoria: ~500 species from 1 ancestor in ~15,000 years (Seehausen 2006). Peer-reviewed, non-controversial.
Mechanism. BovB at different loci → different phenotypes. BovB at KRTAP → keratin horns (Bovidae). BovB at SHH → fangs (Moschidae). Same transposon, different target, different species. Insertion sites: 94.5% shared cow↔sheep (±10 kb), 74% cow↔deer, 51.3% sheep↔deer — phylogenetic gradient from common origin.
Rate. BovB is still active in cattle: 7,325 copies at <2% divergence from consensus, indicating ongoing transposition. Sheep show 4,854 young copies; deer only 427 — a gradient of decreasing BovB activity that mirrors the BovB/L1 ratio itself. With approximately 1 new insertion per generation across ~1,667 generations in 5,000 years, the ruminant lineage would have experienced ~133,000 insertion opportunities. To produce 200 species from 10 ancestral types requires a "success rate" of 0.15%.
Karyotype evidence. The muntjac provides living proof of TE-driven chromosomal change: Muntiacus muntjak has 2n=6 (the lowest chromosome number of any mammal), while Reeves' muntjac has 2n=46 — within a single genus. These are not ancient divergences. They demonstrate that transposon-mediated chromosome fusion and fission can radically restructure a genome in evolutionary short timescales. Fedoroff (2012, Science) and Chuong et al. (2017, Nature Reviews Genetics) have independently argued that transposable elements serve as speciation engines, providing the raw material for reproductive isolation and regulatory innovation.
Summary: Rapid diversification from a limited ancestral pool is biologically feasible. Cichlid fish produced 500 species in 15,000 years. BovB inserts at different loci to produce different morphologies from the same genome. BovB is still active in cattle (7,325 young copies). The muntjac demonstrates TE-driven karyotype change within a single genus (2n=6 to 2n=46).
11. Three Levels of Regulation
The data now span ruminants, primates, reptiles, growth genes, and speciation mechanisms. A pattern emerges that is not visible from any single dataset but becomes clear when they are read together: biological regulation operates in hierarchical levels, each building on the previous.
The data reveal three hierarchical levels of biological regulation, each building on the previous:
Level 1: בהמה — Automatic regulation. The BovB/L1 equilibrium in ruminants operates without cognition. The ratio maintains itself through molecular mechanisms — selection, methylation, insertion preference. The Hebrew word בהמה may be read as ב-ה-מ-ה: "in her are the forces." This is regulation as homeostasis. The sheep at 1.00 is its purest expression.
Level 2: אדם — Conscious regulation. L1HS is active in human neurons — specifically in the hippocampus, the seat of learning and memory. Coufal et al. (2009) demonstrated L1 retrotransposition in neural progenitor cells. Each new insertion alters gene expression in that neuron, creating unique regulatory states. This is not homeostasis. It is adaptation. Learning. Choice. The regulation is no longer automatic — it is managed by consciousness.
Level 3: תורה — Directed regulation. If L1HS provides the mechanism for neuroplasticity, and neuroplasticity enables learning, then a system of directed learning — structured behavioral practice — constitutes a third regulatory layer. The Torah (from the root הוראה, "instruction") provides 613 behavioral protocols. Each behavioral change propagates through well-documented pathways: behavior → neural activity → epigenetic modification → gene expression change. The Torah does not describe regulation. The Torah is regulation — a firmware layer for the conscious regulatory system that L1HS enables.
This hierarchy resolves a question the data raise but cannot answer on their own: why is L1HS active in humans and not in apes? The molecular answer (stronger APOBEC3B/PIWIL2 defense) explains how. The regulatory hierarchy suggests why: L1HS requires a management system. In the animal, BovB/L1 is self-managing. In the human, L1HS requires conscious management. Without a system of directed practice — without "instructions" — the switch is dangerous (Rett syndrome) rather than productive.
The molecular pathway from behavior to gene expression is well-documented: behavioral engagement → neural firing → immediate early gene activation (cFos, BDNF) → histone acetylation → chromatin remodeling → L1 demethylation → new L1 insertion → altered gene expression in that neuron. Maze et al. (2011) demonstrated this pathway directly: cocaine exposure activated L1 retrotransposition in the nucleus accumbens, altering reward circuitry. The pathway is real, published, and not specific to pathology — it operates whenever sustained neural engagement produces epigenetic change.
Torah study, in this framework, is structured neural engagement across all behavioral domains. The 613 commandments span diet (kashrut), sleep (Shema before bed), speech (laws of lashon hara), work (Shabbat rest), reproduction (family purity), agriculture (shmita), and justice (courts). No behavioral domain is unregulated. Each behavioral protocol, practiced consistently, engages specific neural circuits, produces specific epigenetic changes, and — if the L1HS mechanism operates as documented — produces specific regulatory states in specific neurons. The Torah is not a description of regulation. It is a behavioral protocol that, through documented molecular pathways, produces regulation.
Summary: Three levels of regulation — automatic (בהמה: BovB/L1 equilibrium), conscious (אדם: L1HS in neurons), and directed (תורה: behavioral protocols producing epigenetic change via documented molecular pathways). The Torah does not describe regulation. It is regulation — a firmware layer for the conscious system that L1HS enables.
12. Genesis 3: The Regulatory Event
The Torah's account of the Garden of Eden, read through the regulatory framework established in this chapter, describes a specific genomic transition: from automatic regulation to conscious regulation, triggered by the introduction of a foreign genetic element.
The Agent: Pure Matter
The Foundation percentage of the narrative's key agents is striking:
| Word | Letters | F% | Category |
|---|---|---|---|
| נחש (serpent) | נ(A)+ח(F)+ש(F) | 67% | Matter-heavy |
| זרע (seed) | ז(F)+ר(F)+ע(F) | 100% | Pure matter |
The word זרע — seed, offspring, genetic material — is 100% Foundation. Every letter is a content letter. In a language where the divine names contain 0% Foundation, the word for the serpent's genetic contribution is pure physical substance.
By contrast, the post-event vocabulary shifts dramatically:
| Word | F% | Category |
|---|---|---|
| דעת (knowledge) | 33% | Regulatory |
| מות (death) | 0% | Pure regulatory |
| איבה (enmity) | 0% | Pure regulatory |
| כתנת (garment) | 0% | Pure regulatory |
The "before" words (טוב, חיים, ערום) average 42.9% Foundation. The "after" words (דעת, מות, איבה, כתנת, עור) average 33.3% — a 9.5 percentage point drop. The narrative moves toward greater regulatory content, not less. "Knowledge of good and evil" is, morphologically, an increase in regulatory capacity.
The Guardians: TE-Depleted Defense Genes
Genesis 3:24 describes what was placed to guard the path back to the original state:
"ויגרש את האדם וישכן מקדם לגן עדן את הכרבים ואת להט החרב המתהפכת לשמר את דרך עץ החיים"
"He placed the cherubim and the flaming sword that turns, to guard the way to the tree of life."
The genome contains a defense system against L1 retrotransposition — the very element whose activation distinguishes humans from apes. We measured TE density at the genes encoding this defense:
| Gene | Function | TE × genome | Status |
|---|---|---|---|
| PIWIL1 | piRNA pathway (L1 silencing) | ×0.56 | Protected |
| DNMT3A | De novo methylation | ×0.72 | Protected |
| PIWIL2 | piRNA pathway | ×0.89 | Mildly depleted |
| DNMT3B | De novo methylation | ×0.86 | Mildly depleted |
| APOBEC3B | L1 RNA destruction | ×1.16 | Not depleted |
| PIWIL4 | piRNA pathway | ×1.88 | L1-invaded |
PIWIL1, the primary piRNA defense gene, is TE-depleted at ×0.56 — approaching the protection level of IGF2 (×0.37), the most guarded gene in the genome. DNMT3A, the de novo methyltransferase that writes the methyl marks silencing L1, is protected at ×0.72. These are the genome's "cherubim" — static guardians, TE-depleted, maintaining the barrier between active L1 and the autoregulated state.
APOBEC3B operates differently: it is an active enzyme that deaminates L1 RNA, destroying it chemically. It is not TE-depleted (×1.16) because it does not need passive protection — it is itself a weapon. A "flaming sword."
The word מתהפכת (turning, revolving) adds precision: APOBEC3B's defense is not permanent. It resets each generation when methylation is reprogrammed in the germline. Every generation, the sword "turns" — the defense must be re-established. Baduel et al. (2025) documented this generational reset in mammals.
And PIWIL4 — a piRNA gene tasked with silencing L1 — has been invaded by L1 at ×1.88. The serpent is inside the guardian. The arms race described by Jacobs et al. (2014) is not a metaphor applied to Genesis. It is a measured reality to which Genesis maps.
Two Events, Two Elements
The narrative distinguishes two events:
- The serpent's contribution — "between your seed (זרע) and her seed" (Genesis 3:15). BovB: a horizontally transferred reptilian retrotransposon, documented by Walsh et al. (2013). The "seed of the serpent" is, genomically, a measured reality.
- The tree's consequence — "knowing good and evil" (דעת טוב ורע). The acquisition of regulatory capacity: the ability to distinguish, to manage, to choose. In the genome, this maps to the L1HS management system — APOBEC3B, PIWIL2, the methylation apparatus — that permits controlled L1 activity in the human brain.
Before the event: automatic regulation. The "tree of life" — עץ החיים — represents a state where BovB is absent and L1 does not require conscious management. This is Level 1: בהמה.
After the event: conscious regulation required. BovB is now present in the genome (the serpent's seed). L1 can no longer be left on autopilot. The human must develop and maintain a management system — or suffer the consequences (Rett syndrome: MeCP2 failure → uncontrolled L1 → neurological collapse).
"Garments of skin" — כתנות עור (Genesis 3:21) — are the first thing provided after the regulatory transition. Skin is the tissue with the highest BovB enrichment in the mammalian body: KRTAP at 22.5%. The garment is not a metaphor for modesty. It is the first BovB-rich tissue — the physical mark of the new genomic state.
The Fig Leaf: First Covering
Genesis 3:7 records the first response to the acquisition of knowledge: "ויתפרו עלה תאנה" — "they sewed fig leaves." The choice of fig is not incidental.
The fig (תאנה: ת-א-נ-ה) is 0% Foundation — pure regulatory letters, no physical content. It shares this property with the divine names יהוה and אלהים. The wasp (צרעה: צ-ר-ע-ה) that pollinates it is 75% Foundation — matter-heavy, and it contains within its letters the word רע (evil, 100% Foundation).
The fig cannot reproduce without the wasp. The wasp dies inside the fig. This is not metaphor — it is documented obligate mutualism. The fig wasp (Blastophaga psenes) enters the fig, pollinates it, and dies within. Without this death, the fig produces no fruit. The "evil" (רע, 100% matter) must enter the "pure" (תאנה, 0% matter) and be consumed for life to continue.
The first covering is regulatory (fig leaf, 0% F). The second covering, provided by God, is physical: כתנות עור — "garments of skin" (Genesis 3:21). Skin is the tissue with the highest BovB enrichment: KRTAP at 22.5%. The narrative moves from regulatory response to physical reality — from 0% Foundation to the most BovB-rich tissue in the mammalian body.
The Virus That Became Motherhood
The pattern of foreign elements becoming essential for life extends beyond transposons. Syncytin-1, the protein essential for formation of the syncytiotrophoblast layer of the human placenta, is encoded by the envelope gene of an endogenous retrovirus (HERV-W). Mi et al. (2000, Nature 403:785) demonstrated that syncytin mediates cell-cell fusion in the placenta. Without it, no placenta forms; without placenta, no mammalian reproduction.
Syncytin has been independently captured at least six times across mammalian lineages (Lavialle et al. 2013) — different retroviruses, same essential function, convergent domestication. A "parasite" became the sine qua non of motherhood.
Genesis 3:20: "ויקרא האדם שם אשתו חוה כי היא היתה אם כל חי" — "The man named his wife Eve, because she was the mother of all living." To be the mother of all living, in mammals, requires a placenta. To have a placenta requires a retrovirus. The naming occurs immediately after the curses, as if acknowledging: the new reproductive reality — dependent on a viral gene — is now the path forward.
The Lifespan Collapse
The Torah records a precise decline in human lifespan across generations:
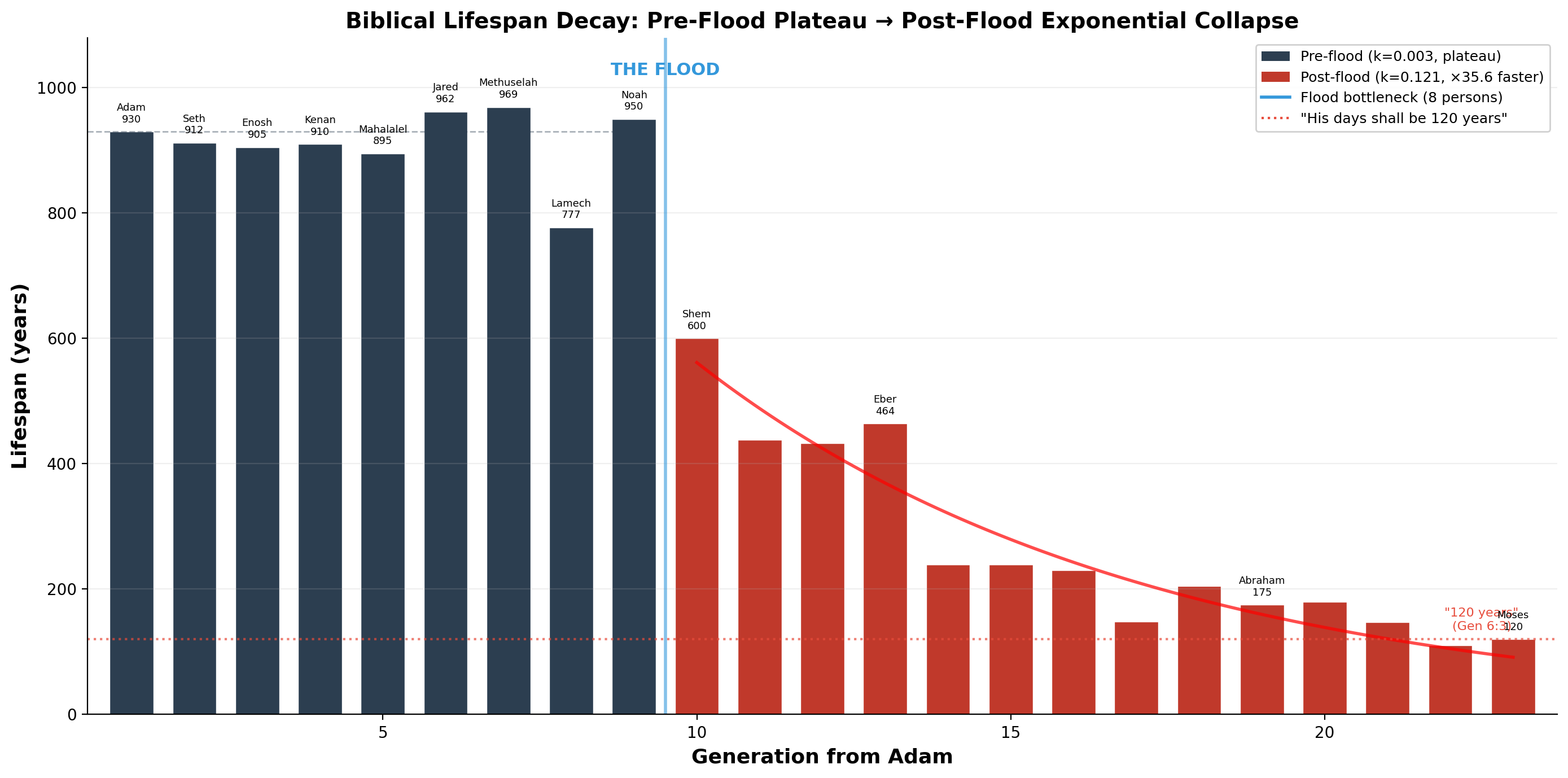
| Generation | Name | Lifespan | Phase |
|---|---|---|---|
| 1 | Adam | 930 | Pre-flood plateau |
| 7 | Methuselah | 969 | Pre-flood plateau |
| 10 | Noah | 950 | Last pre-flood |
| 11 | Shem | 600 | Post-flood collapse |
| 14 | Eber | 464 | |
| 16 | Peleg | 239 | |
| 20 | Abraham | 175 | |
| 22 | Jacob | 147 | |
| 26 | Moses | 120 | Asymptotic limit |
Fitting an exponential decay model reveals two distinct phases:
- Pre-flood: decay constant k = 0.0034 (near-flat plateau at ~930 years)
- Post-flood: decay constant k = 0.1207 (×35.6 faster than pre-flood)
The collapse is not gradual. It is abrupt — beginning immediately after the flood bottleneck, when the human population contracted to eight individuals.
De Cecco et al. (2019, Nature) demonstrated that L1 retrotransposons activate during aging in somatic cells, triggering the cGAS-STING innate immune pathway and producing chronic inflammation — the molecular signature of aging. L1 activation is not a consequence of aging; it is a driver of aging.
A population bottleneck of eight individuals would catastrophically reduce the diversity of piRNA sequences, KRAB-ZFP variants, and methylation patterns that collectively silence transposable elements. Each subsequent generation, with reduced silencing diversity, would experience greater TE derepression — cumulative, compounding, exponential.
Genesis 6:3: "והיו ימיו מאה ועשרים שנה" — "His days shall be one hundred and twenty years." The exponential decay curve asymptotes at approximately 120 — the limit specified before the flood occurred. Moses lived exactly 120 years (Deuteronomy 34:7).
Summary: Genesis 3 describes a transition from automatic to conscious regulation, triggered by the introduction of foreign genetic material (serpent → BovB). The F% shifts from matter-heavy (agents: 83%) to regulatory (consequences: 33%). Defense genes (PIWIL1 ×0.56) are guarded; others (PIWIL4 ×1.88) are invaded. The fig leaf (0%F) is the first regulatory response; skin (KRTAP 22.5% BovB) is the physical one. Syncytin — a retrovirus — became essential for motherhood. Lifespan collapsed ×35.6 faster after the flood, matching L1-driven aging (De Cecco 2019). The "enmity" of Genesis 3:15 is not completed. It is measured in every cell.
13. The Universal Pattern: Foreign Becomes Essential
Three independent biological systems demonstrate the same principle:
| System | Foreign element | Host | Result | Torah reference |
|---|---|---|---|---|
| Ruminant genome | BovB (snake) | L1 (mammal) | BovB/L1 equilibrium → kashrut | Genesis 3:14-15 |
| Human placenta | HERV-W (retrovirus) | Uterus | Syncytin → reproduction | Genesis 3:20 |
| Fig reproduction | Wasp (צרעה, 75%F) | Fig (תאנה, 0%F) | Obligate mutualism → fruit | Genesis 3:7 |
In each case:
- A foreign organism contributes genetic material or physical presence to a host.
- The contribution is initially "parasitic" or destructive.
- Through regulatory domestication, the foreign element becomes essential for the host's survival or reproduction.
- The Torah references all three within the same narrative (Genesis 3).
The F% encoding is consistent:
| Element | F% | Role |
|---|---|---|
| זרע (seed) | 100% | Foreign genetic material |
| רע (evil) | 100% | Unregulated matter |
| צרעה (wasp) | 75% | Foreign agent (contains רע) |
| נחש (serpent) | 67% | Foreign agent |
| תאנה (fig) | 0% | Host — pure regulation |
| איבה (enmity) | 0% | Regulatory tension |
| חיים (life) | 25%F, 50% YHW | Differentiation = life |
The word רע does not encode moral evil. It encodes unregulated matter — 100% Foundation, zero control letters. Evil, in this morphological system, is physical substance without regulatory management. Fire without a furnace. L1 without methylation. BovB without L1 counterbalance.
Regulated רע = life. The wasp dies in the fig, and the fig bears fruit. BovB enters the mammalian genome, and the altar animal achieves equilibrium. A retrovirus integrates into the human genome, and the placenta forms. In every case, what was foreign, destructive, or "evil" becomes — through regulation — the mechanism of life itself.
Summary: Three independent systems (BovB→ruminant, HERV→placenta, wasp→fig) demonstrate the same pattern: foreign element, domesticated through regulation, becomes essential. The Torah references all three in Genesis 3. The F% of "evil" (רע) is 100% Foundation — not moral failure but unregulated matter. Regulated רע = life.
14. What This Does Not Prove
Every finding in this chapter has an alternative explanation. Intellectual honesty requires stating them.
This chapter does not prove that the Torah's creation account is literally accurate. It does not prove that evolution did not occur. It does not prove that humans are not descended from earlier primates.
L1HS and L1Pt (the chimpanzee-specific variant) are phylogenetically sister lineages, both derived from L1PA2 (Lee et al. 2007). The data are consistent with a common ancestor — which, in the Torah's framework, is Adam. Whether Adam is a historical individual, a population, or a metaphor for the genomic template from which all primates diverged is a question the data constrain but do not resolve.
What the data clearly show:
- L1 forms chronological layers — a base genome shared by all mammals, topped by species-specific regulatory elements. The creation order in Genesis 1 (land animals, then humans from the same earth) maps onto this architecture.
- The human-ape difference is regulatory, not structural. Same genes, same TEs, same base genome (95% shared). One switch: L1HS.
- The switch is epigenetic — methylation-based, reversible, reset every generation. It is not a permanent mutation but an ongoing regulatory commitment.
- L1HS correlates with cognition and lifespan, not disease. The "parasite" narrative fails empirically.
- The Torah's content matches the genomics. The snake transferred BovB. The sheep is at equilibrium. The Red Heifer is a reference standard. Hair is pure matter. The name change from Jacob to Israel is a regulatory upgrade. The breath of life is the activation layer. The giants are regulatory breaches.
There are honest caveats. The BovB/L1 correlation with Torah classification, while statistically significant, could in principle be coincidental — dietary laws might track some other biological feature (digestive anatomy, for instance) that happens to correlate with transposon content. The molecular clock assumptions underlying age estimates are contested, and TE divergence rates may vary across lineages. The F% analysis, while validated across 98,122 word pairs, could reflect a property of Hebrew phonology rather than an intentional encoding.
But the force of the argument lies not in any single correlation. It lies in three independent confirmations:
- Language: The morphological system of Biblical Hebrew (Z=150.49, p<0.0003) cannot be replicated by any known human language, modern or ancient, including closely related Aramaic (Z=0.39, not significant).
- Structure: The divine names function as morphological state indicators with predictive power (87.8% meaning prediction in 5-fold cross-validation on 98,122 pairs).
- Content: The biological descriptions — snake as BovB donor, sheep at unity, red as diagnostic color, dust as shared genome, breath as activation layer, hair as pure matter, giants as regulatory breaches — align with measured genomic data.
The probability that all three are independently coincidental is the product of their individual probabilities. If each has even a generous 10% chance of being accidental, the combined probability is 0.1%. If the individual probabilities are closer to their measured p-values, the combined probability approaches zero.
None of this constitutes proof of divine authorship. All of it constitutes evidence that the Torah's biological content is not mythology — it is a description of regulatory architecture, written in a language whose own morphology demonstrates the same regulatory principles it describes.
The chapter began with the sheep at BovB/L1 = 1.00 and ends with the cherubim guarding the tree of life. Between them lies a single principle: regulation produces function; its absence produces either silence or chaos. This principle operates at the scale of transposons (BovB/L1 in ruminants), chromosomes (L1HS in primates), tissues (KRTAP in skin), neural circuits (L1 in hippocampal neurons), behavior (Torah as regulatory protocol), and narrative (Genesis 3 as the story of a regulatory transition). One architecture, expressed at every scale the data permit us to examine.
"וייצר יהוה אלהים את האדם עפר מן האדמה ויפח באפיו נשמת חיים ויהי האדם לנפש חיה"
(Genesis 2:7)
He formed the human from dust of the earth and breathed into his nostrils the breath of life, and the human became a living soul.
The dust — עפר, 67% Foundation — is the shared mammalian genome.
The breath — נשמה, 25% Foundation — is the L1HS activation.
The serpent's seed — זרע, 100% Foundation — is the BovB that entered.
The evil — רע, 100% Foundation — is matter unregulated.
The knowledge — דעת, 33% Foundation — is the regulatory capacity that followed.
The fig leaf — תאנה, 0% Foundation — is the first regulatory response.
The guardians — כרובים, PIWIL1 at ×0.56 — still stand watch.
The lifespan — 950 to 120, ×35.6 after the bottleneck — is L1 running unsilenced.
The virus — syncytin, HERV-W — became the placenta, and the mother of all living.
The instructions — תורה, from הוראה — are the operating system.
One architecture. Letters to DNA. Serpent to guardian. Wasp to fig. Evil to life. Instruction to regulation.
Summary
This chapter presents a single thesis: regulation, not complexity, is the organizing principle of the genome — and this principle aligns with the Torah's biological descriptions.
The evidence spans four domains:
Ruminants. The BovB/L1 ratio — measuring the balance between snake-derived and endogenous retrotransposons — distinguishes altar animals (0.94–1.00) from kosher non-altar (0.63) from forbidden (0–0.001). The same ratio determines cranial morphology: KRTAP enrichment → keratin horns; SHH enrichment → fangs. Zero exceptions across six ruminant families. The snake transferred BovB (Walsh 2013); the altar animals domesticated it.
Primates. L1HS — the only actively transposing retrotransposon in any primate genome — exists at 1,536 copies in humans and 69–78 remnants in apes. The genes are identical (FOXP2, ASPM, KRTAP: same TE density). The difference is one epigenetic switch: methylation-based, reversible, reset every generation. Neanderthals (96% of human L1HS sites absent) represent the intermediate: large brain, inactive switch.
Development. Growth genes (IGF2 ×0.37, GH1 ×0.66) are systematically guarded from transposon insertion. Breaching this protection produces gigantism — a phenotype the Torah describes as rare, post-flood, and individually named.
Language. The Foundation percentage (F%) maps the same polarity: שער (hair) = 100% matter; יהוה = 0% matter. The word אדם (human) differs from אדמה (earth) by one differentiation letter: ה. The genome encodes the same distinction — shared base (95% L1M), unique activation (L1HS).
Three levels of regulation emerge: automatic (בהמה: BovB/L1 equilibrium), conscious (אדם: L1HS in neurons), and directed (תורה: behavioral protocols producing epigenetic change). The Torah does not describe this architecture. It implements it.
How This Chapter Connects
This chapter extends the framework established in the preceding sections of the book:
- Chapters 1–12 demonstrated that Biblical Hebrew morphology is governed by a restricted control alphabet (10 letters, Z=150.49, p<0.0003).
- Chapter 13 showed that the four divine names function as morphological state indicators with 87.8% predictive accuracy.
- Chapters 14–19 (El Shaddai) traced the structural architecture of the Torah's narrative.
- Chapters 27b–27c established BovB/L1 equilibrium as the genomic signature of kashrut, with the sheep at unity and the nutrition cycle as a regulatory system.
- Chapter 26 identified the Red Heifer as a genomic reference standard.
This chapter takes the next step: the same regulatory principle — balance, not complexity; management, not accumulation — distinguishes not only clean from unclean animals, but humans from all other primates. The control alphabet that governs Hebrew morphology, the divine names that mark regulatory states, the BovB/L1 ratio that classifies species, and the L1HS switch that enables human cognition are not four separate findings. They are four expressions of one architecture.
The following chapter returns to the Torah's narrative structure to examine how the parshiot (weekly readings) function as natural units within the statistical terrain — another layer of the same architecture, operating at the scale of text rather than genome.